Human Cancer Signatures
Version 3.5Single Base Substitutions (SBS) Signatures
About the collection
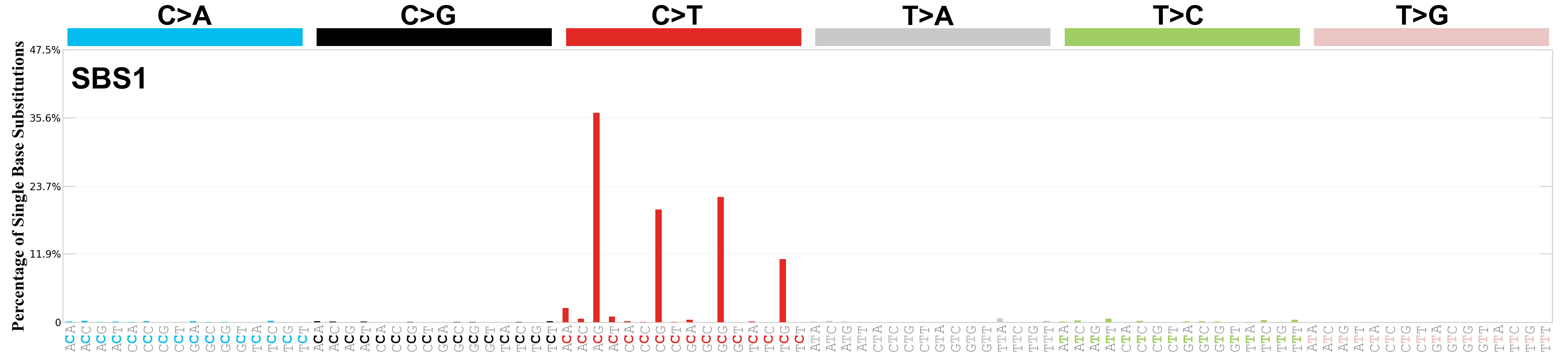
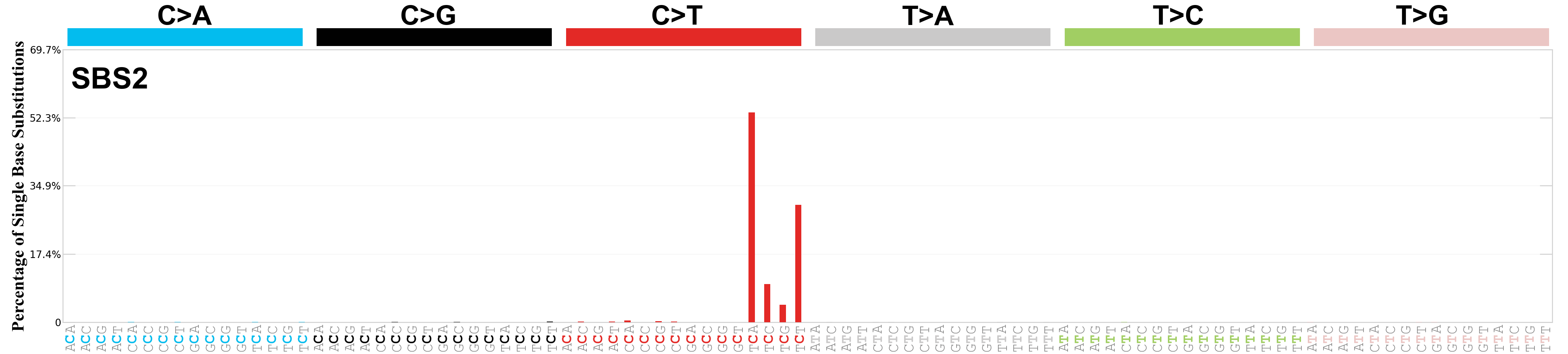
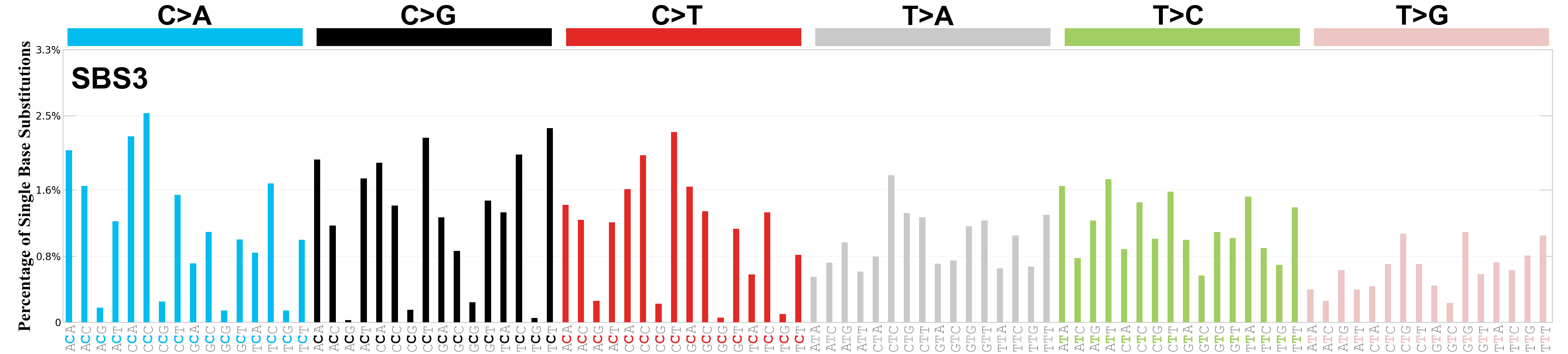
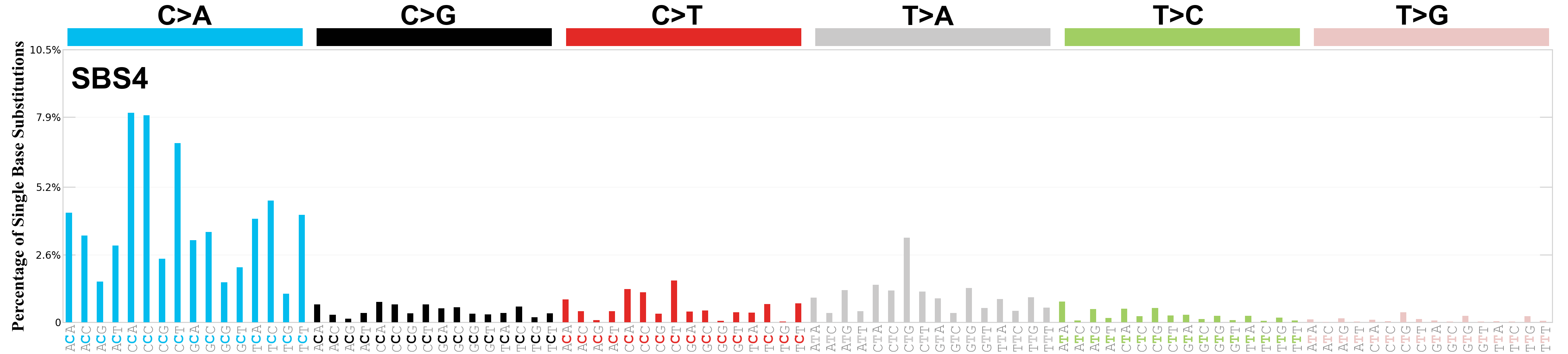
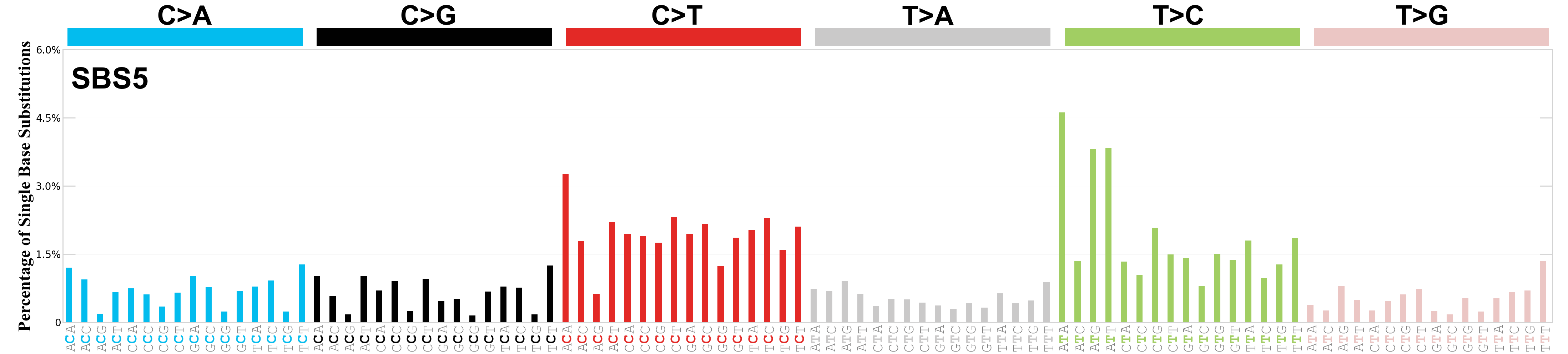
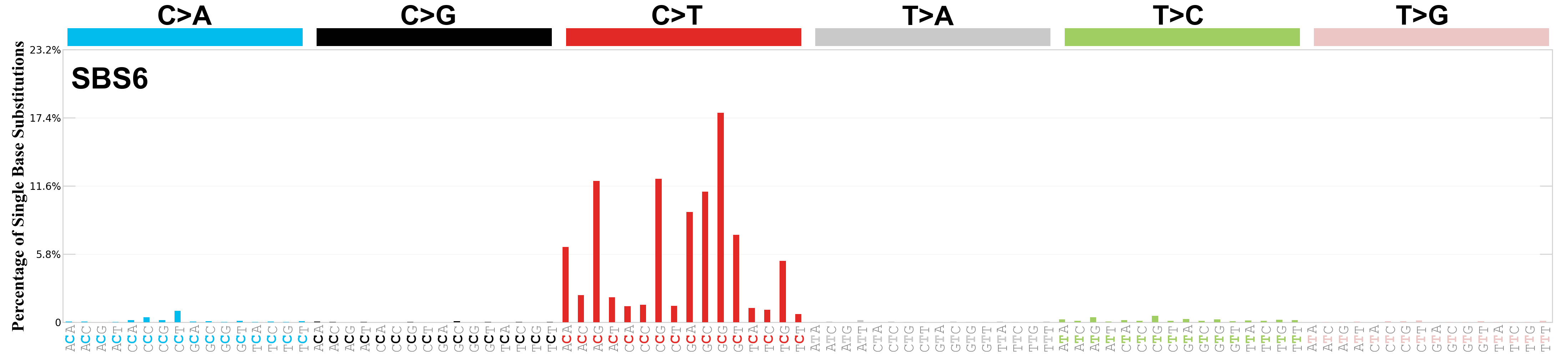
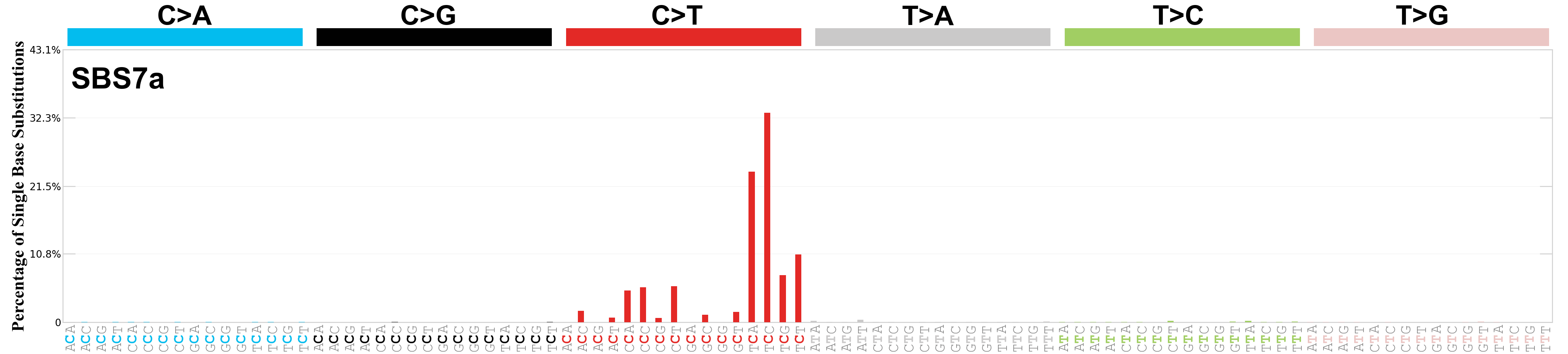
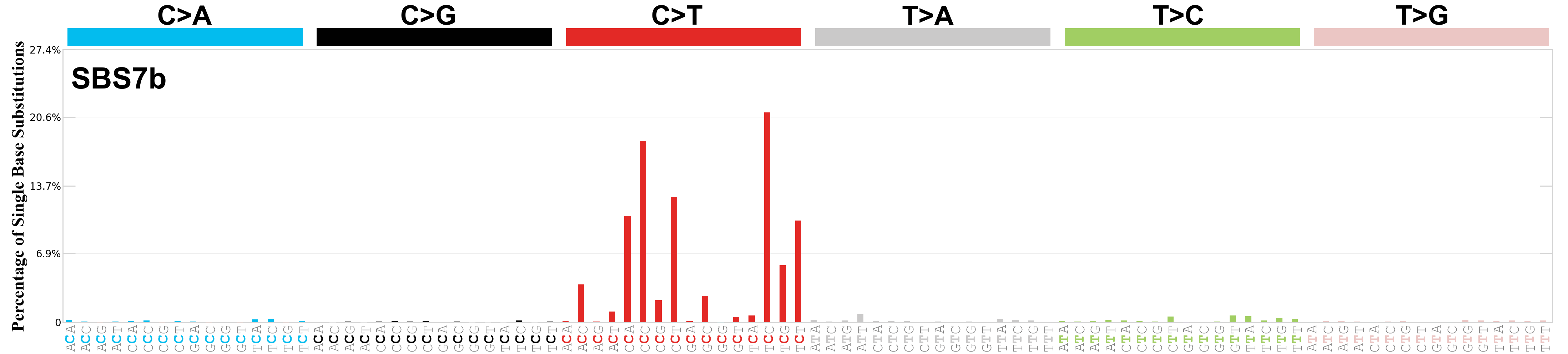
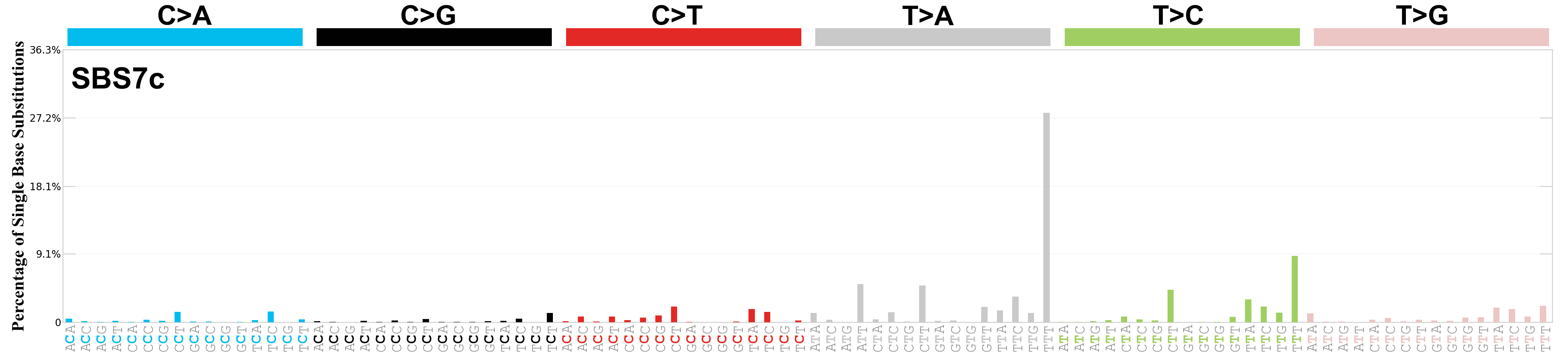
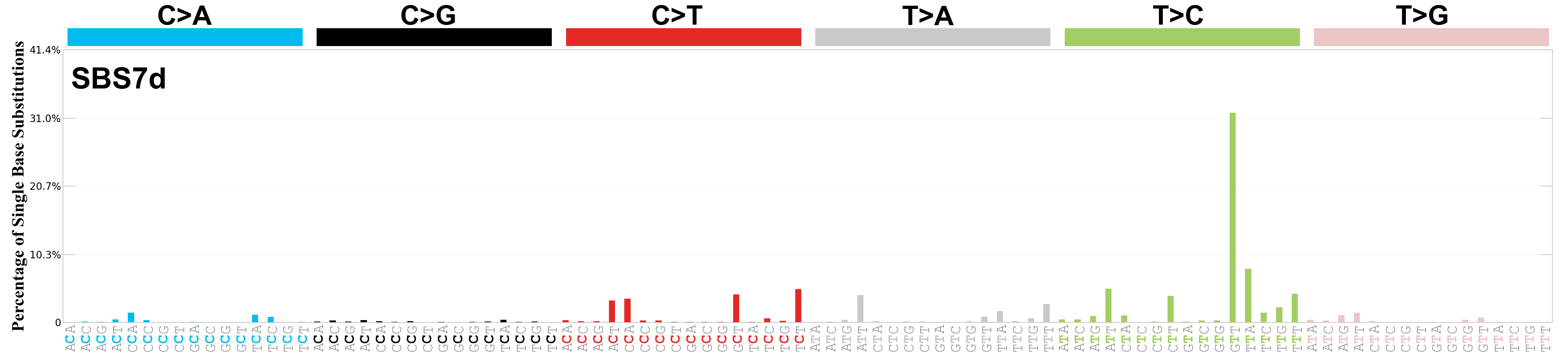
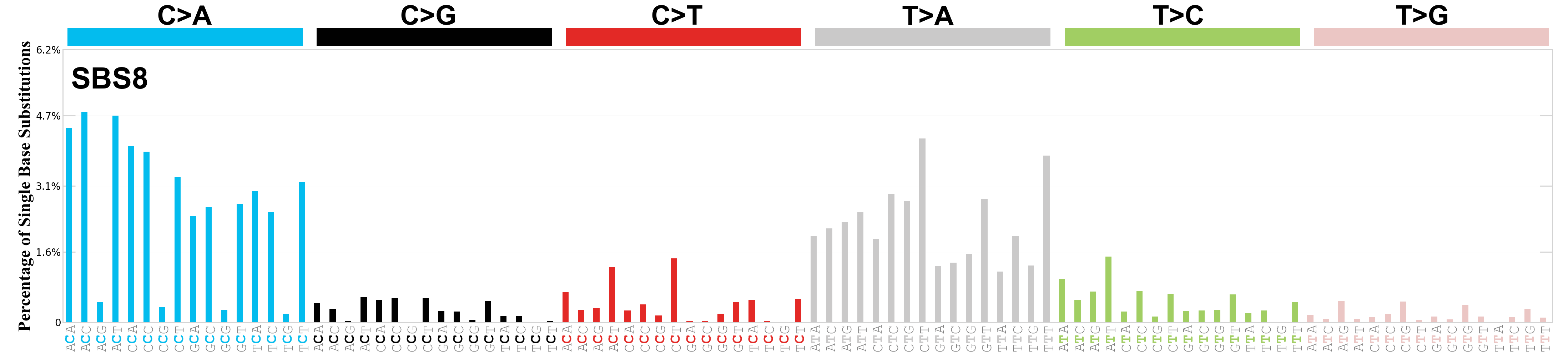
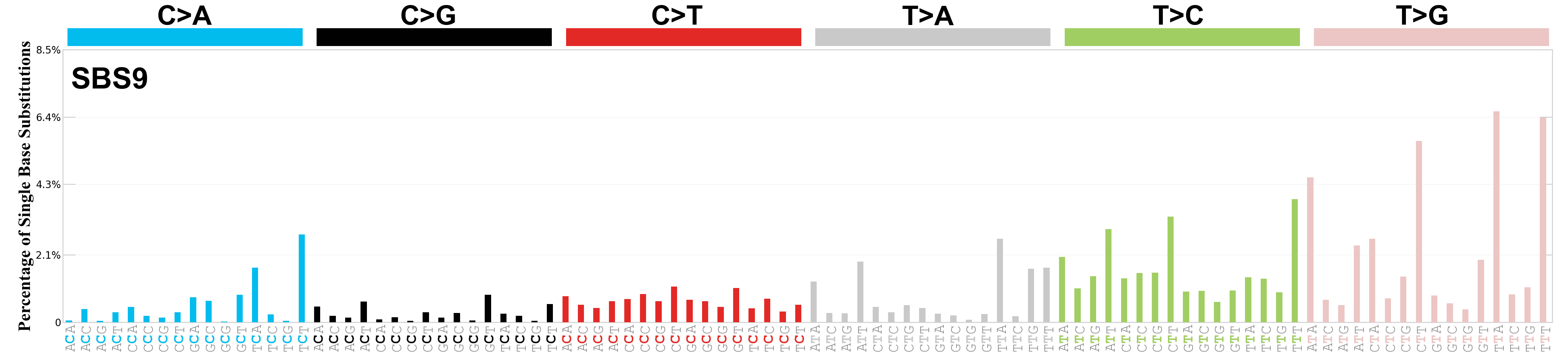
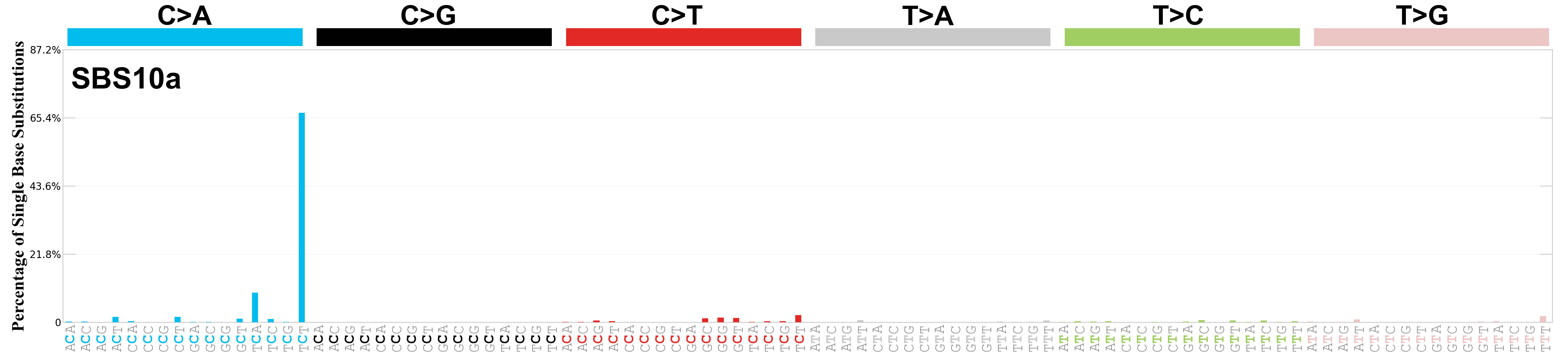
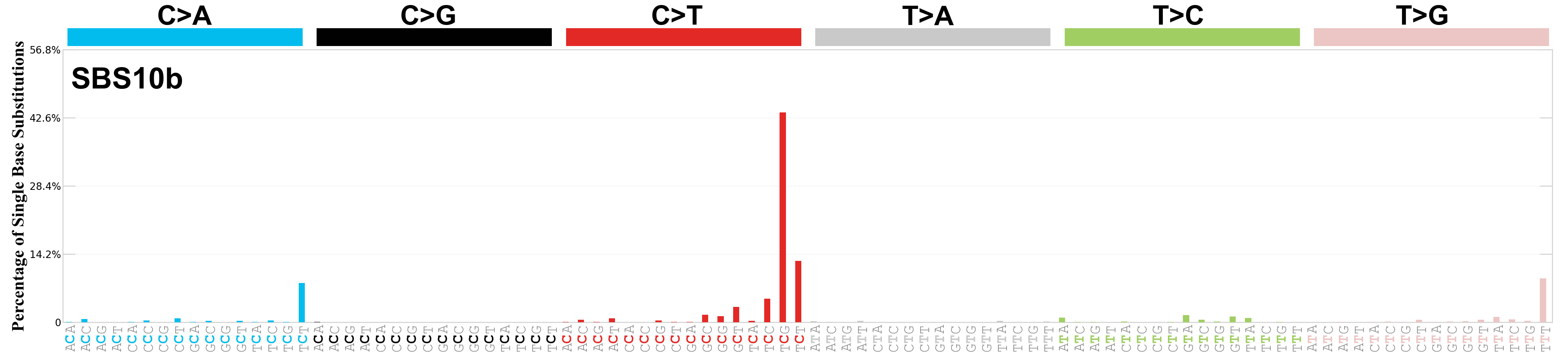
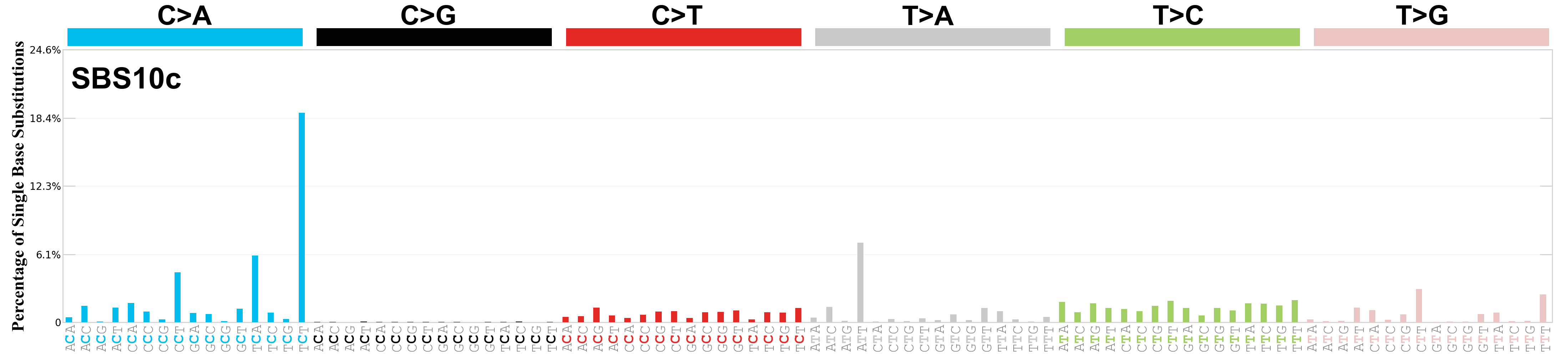
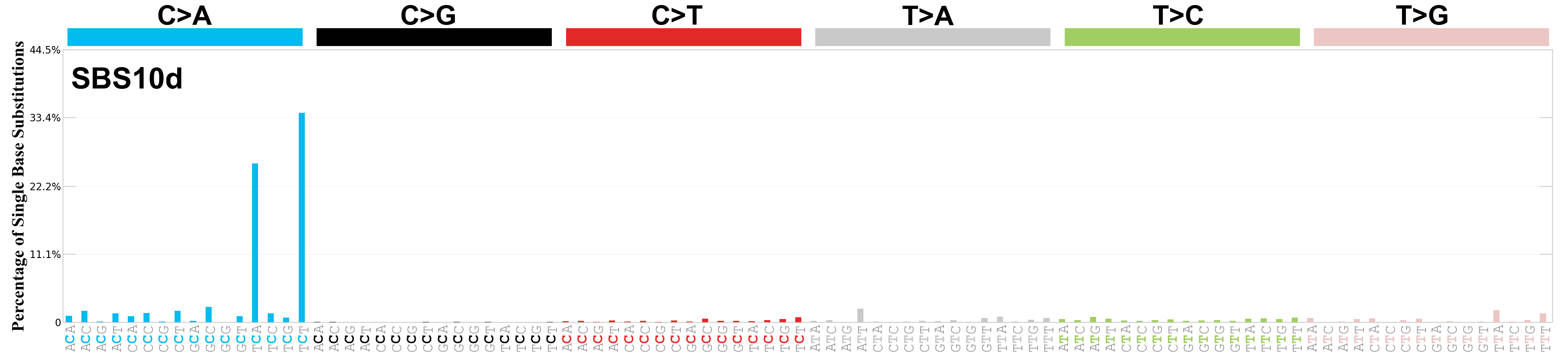
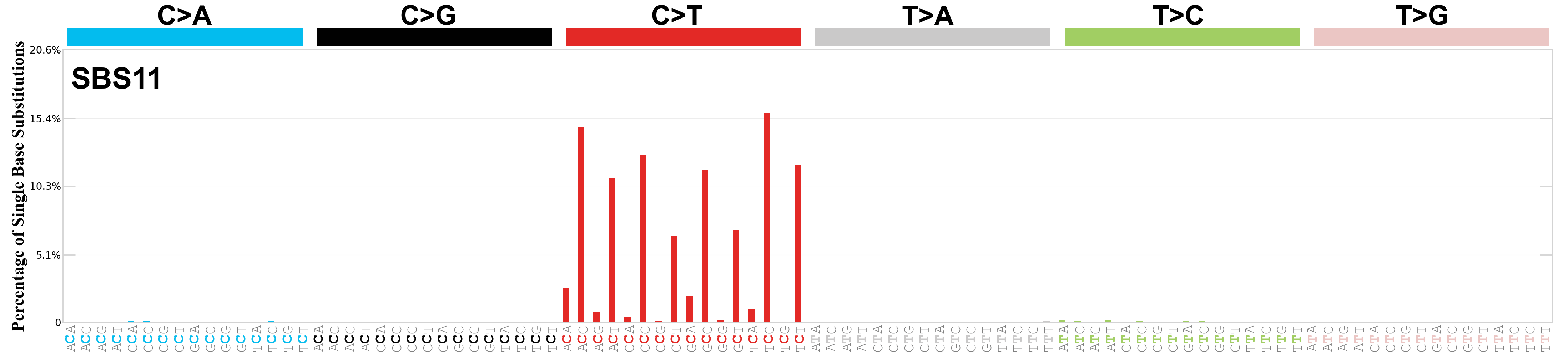
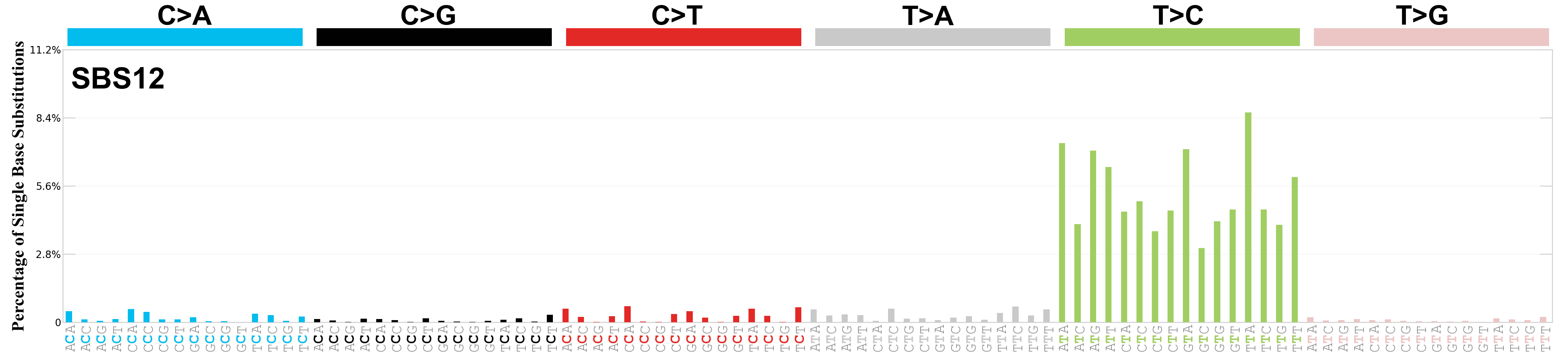
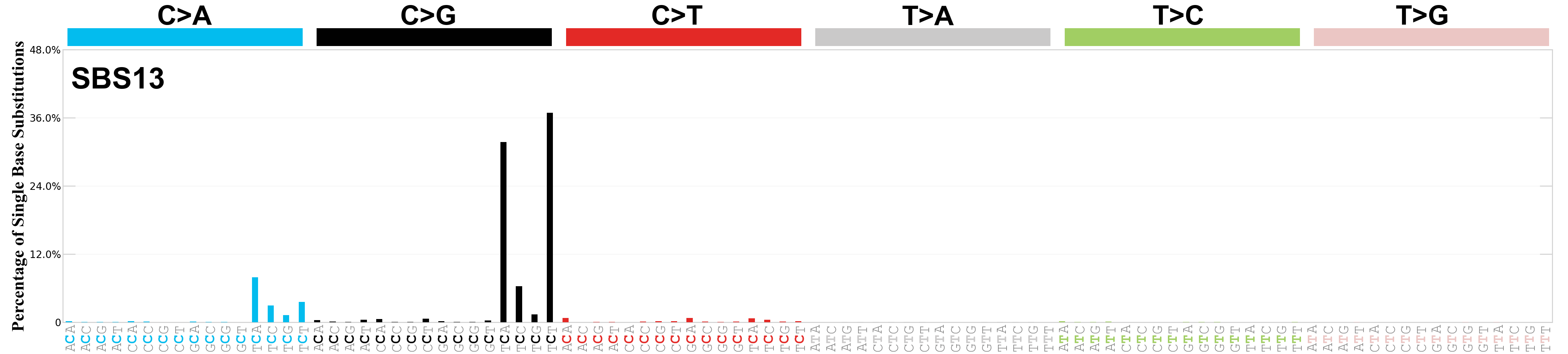
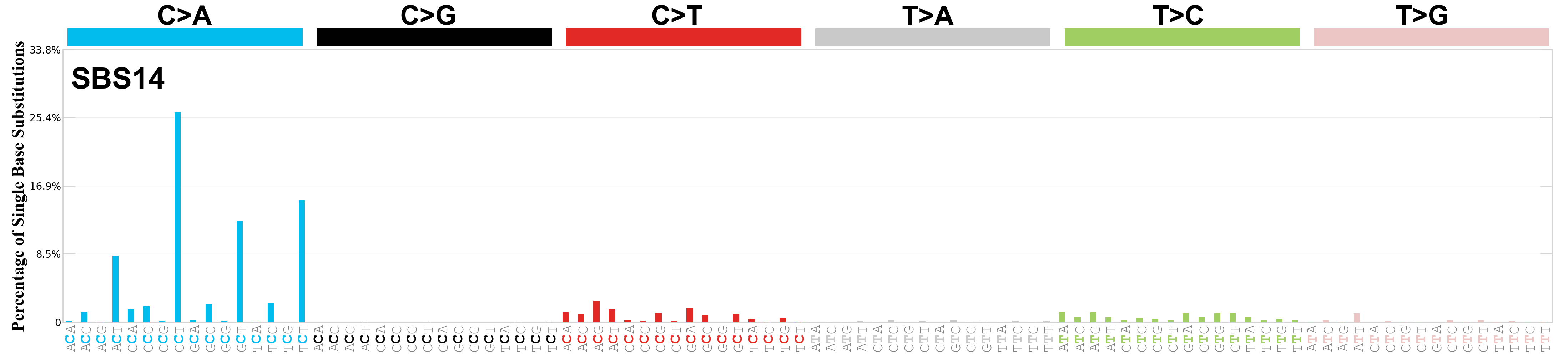
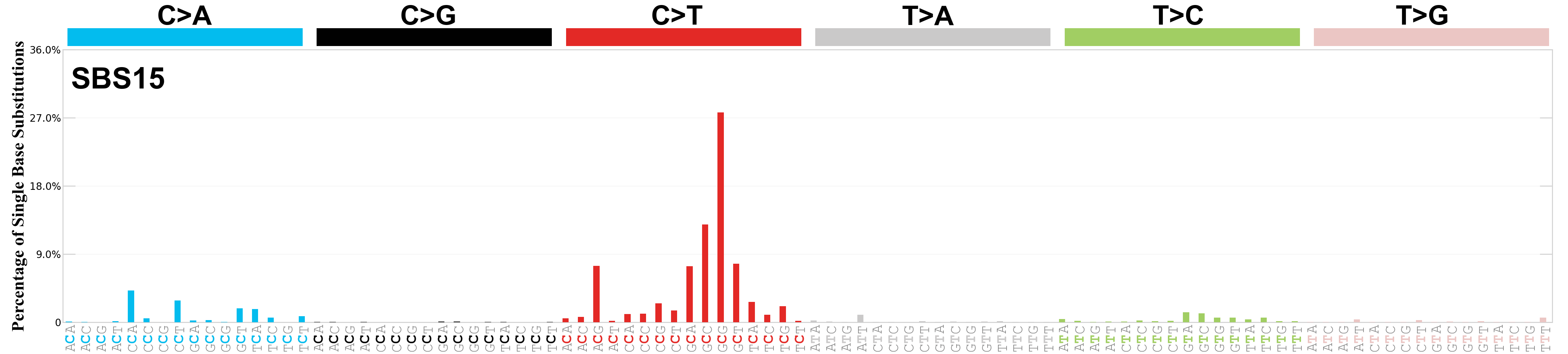
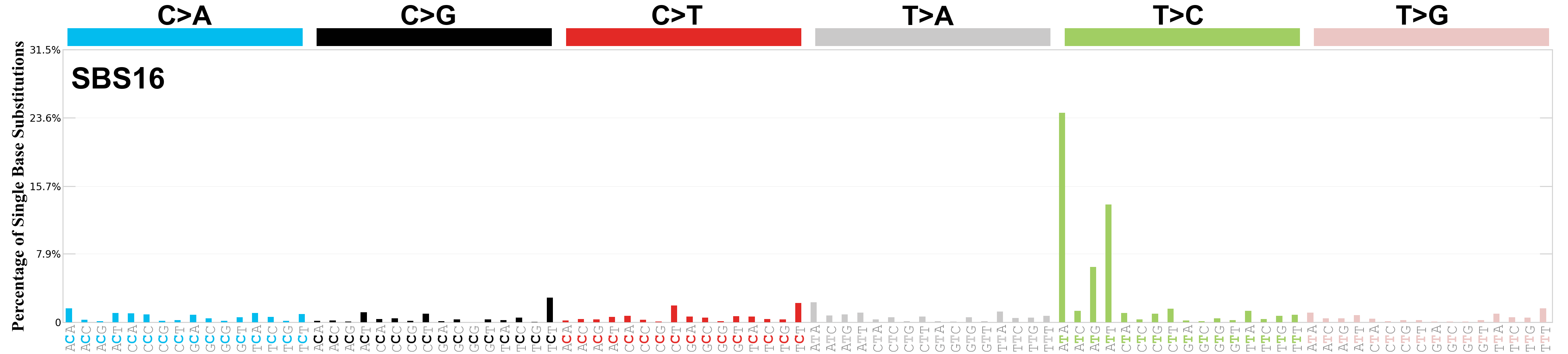
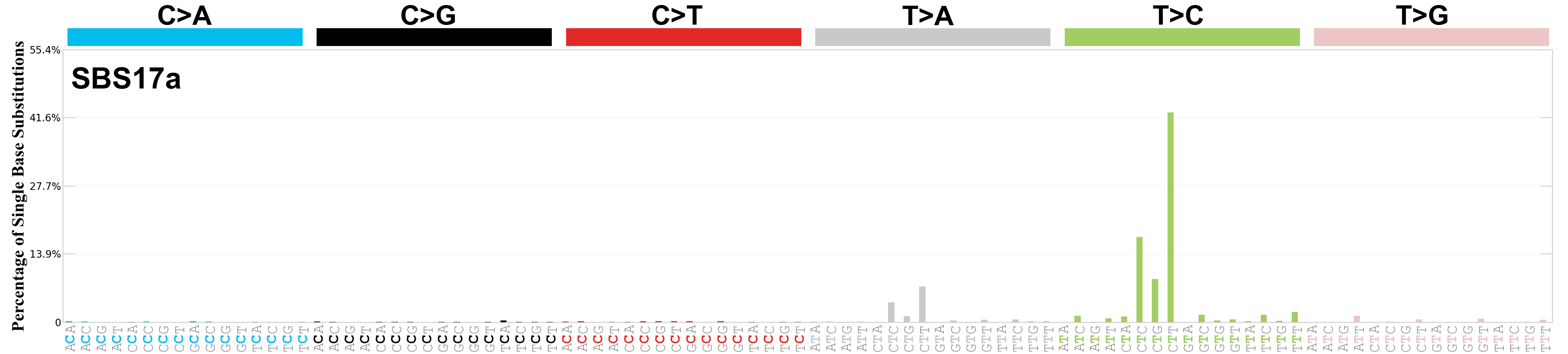
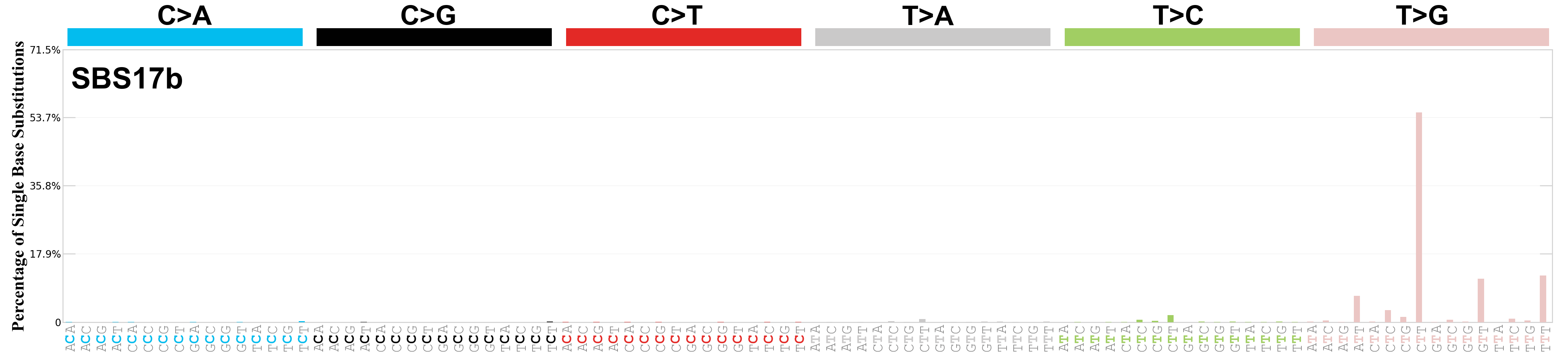
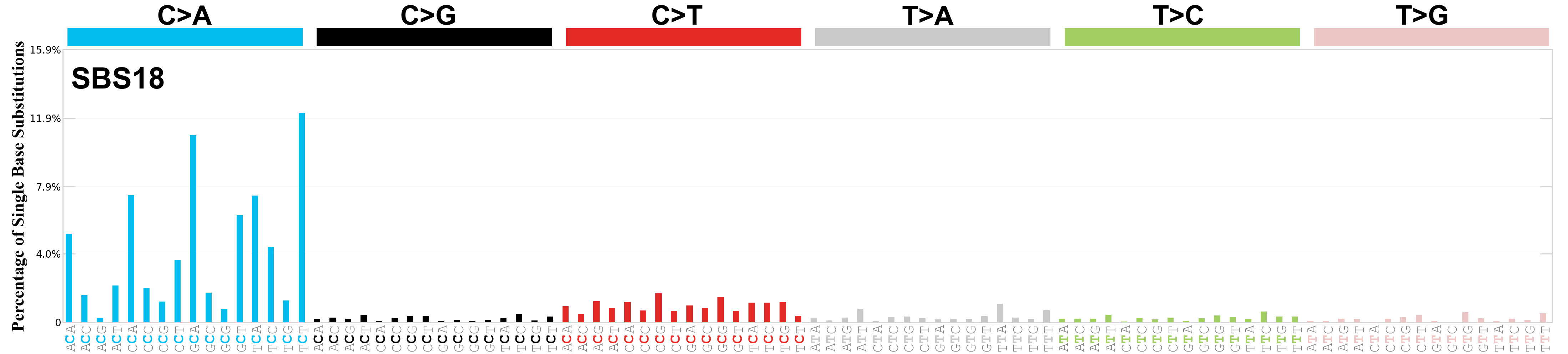
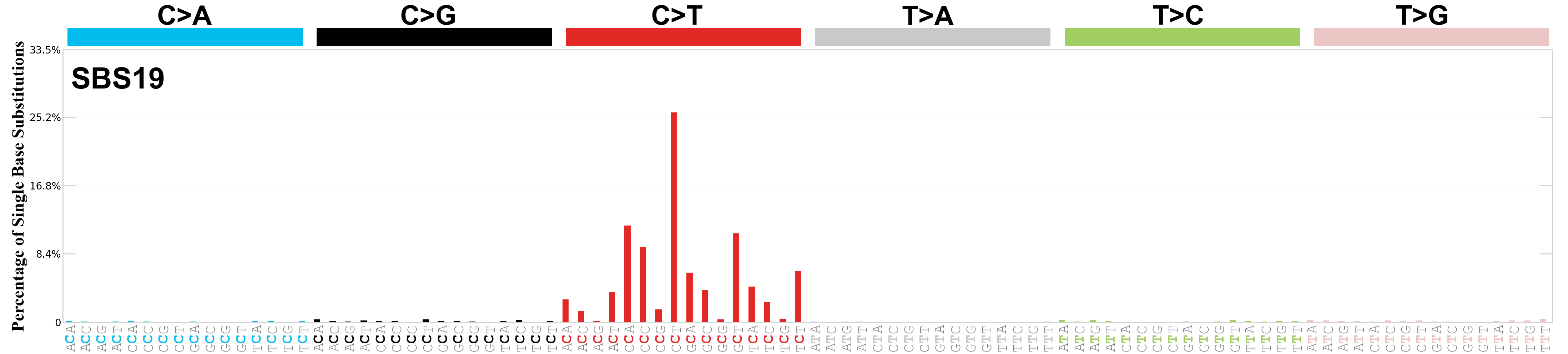
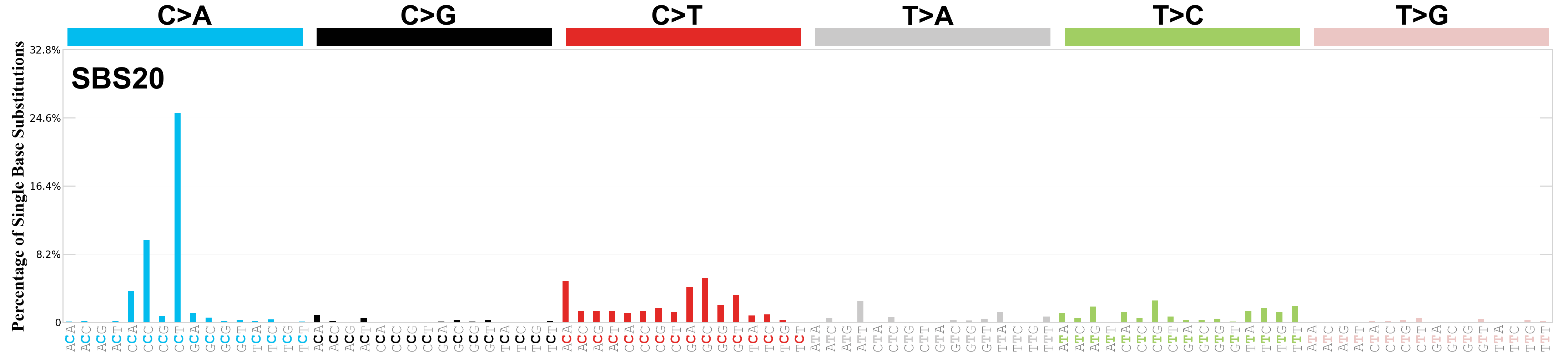
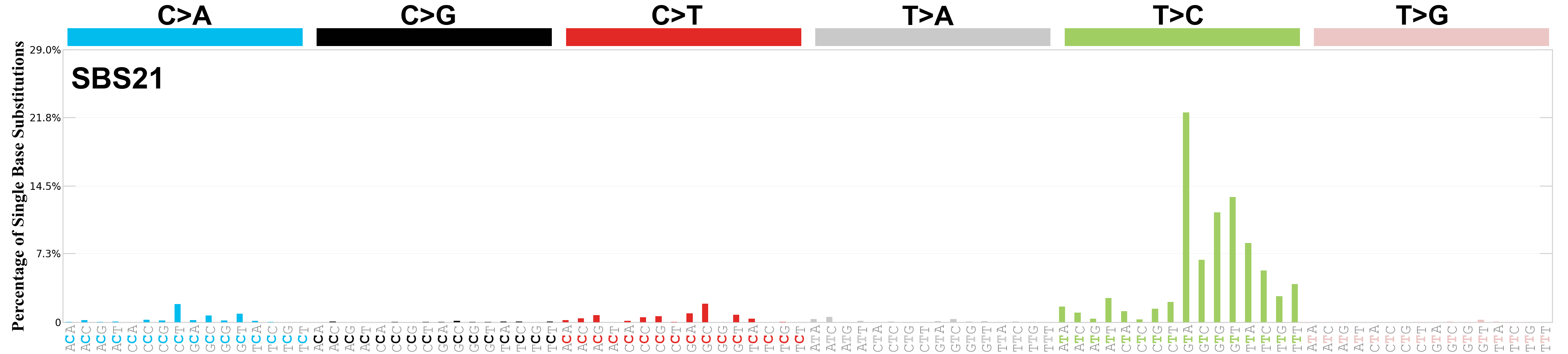
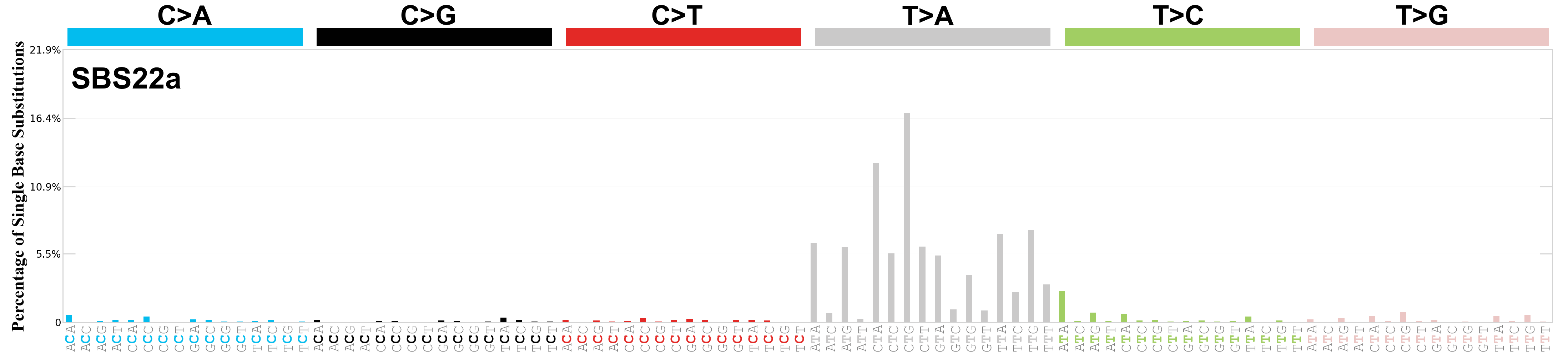
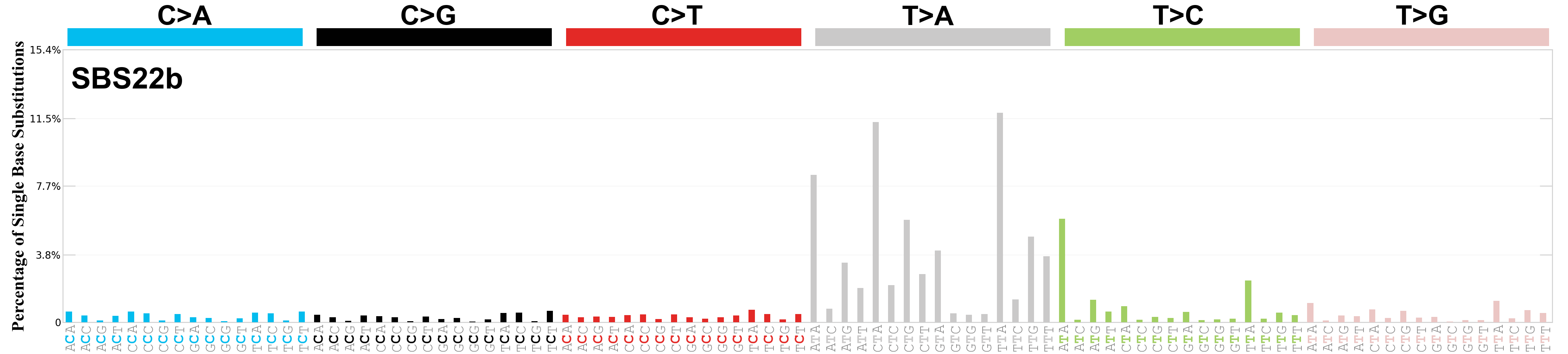
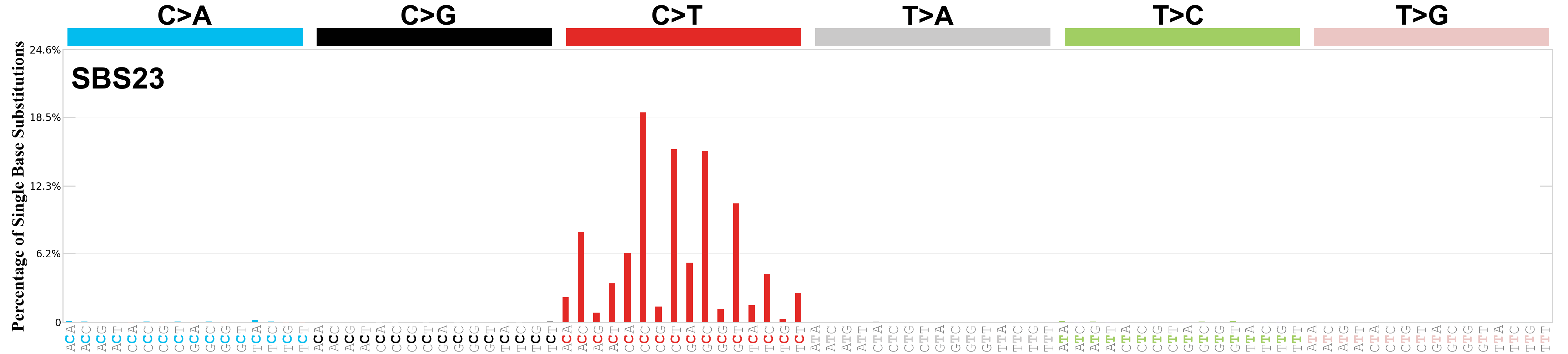
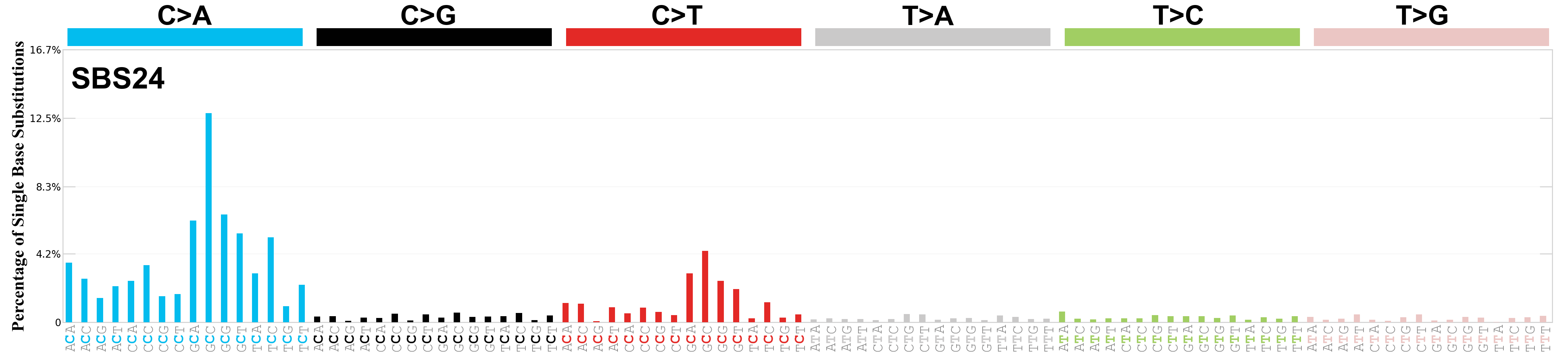
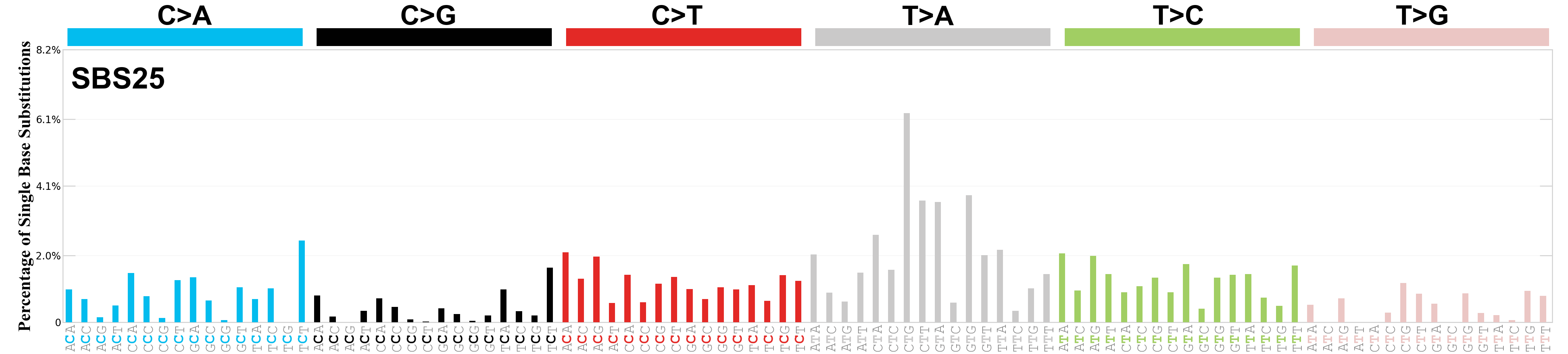
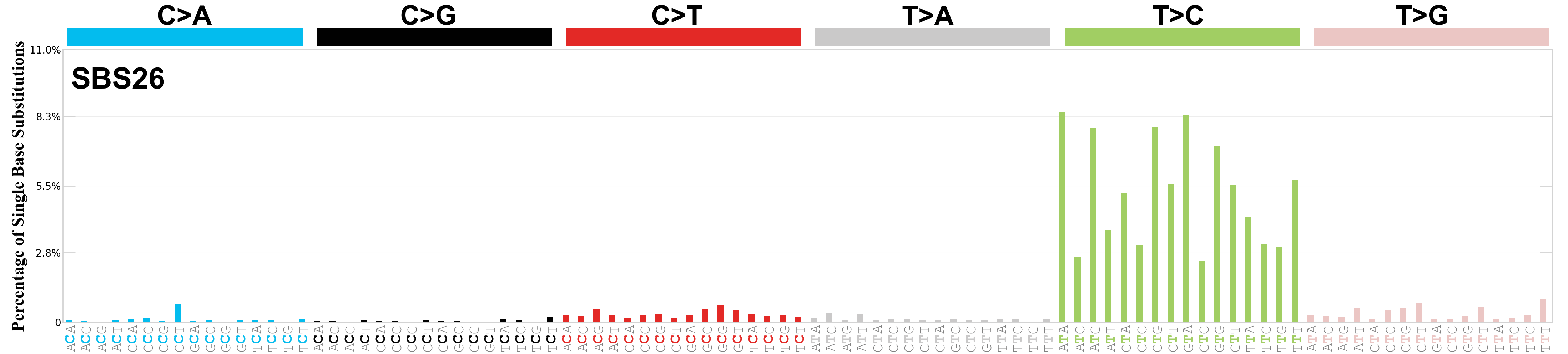
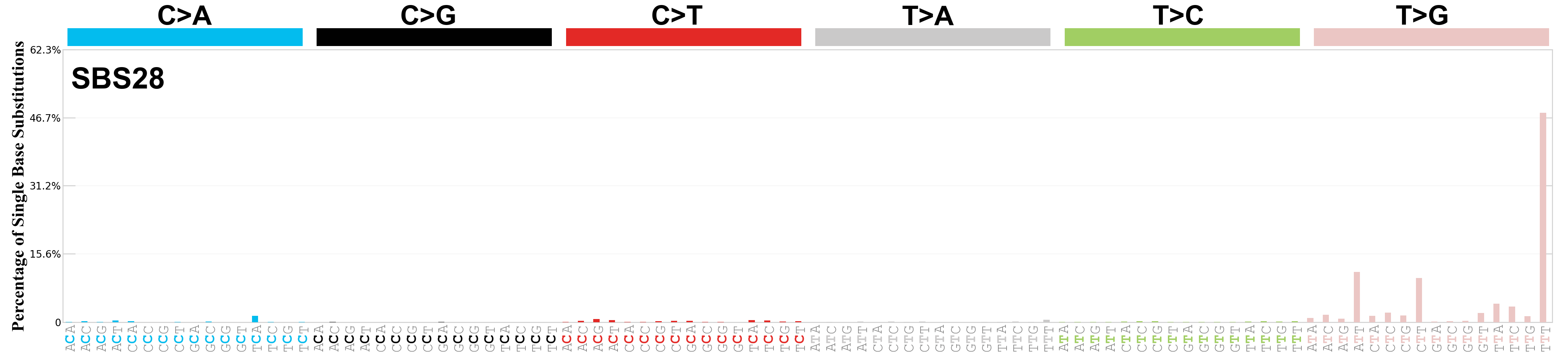
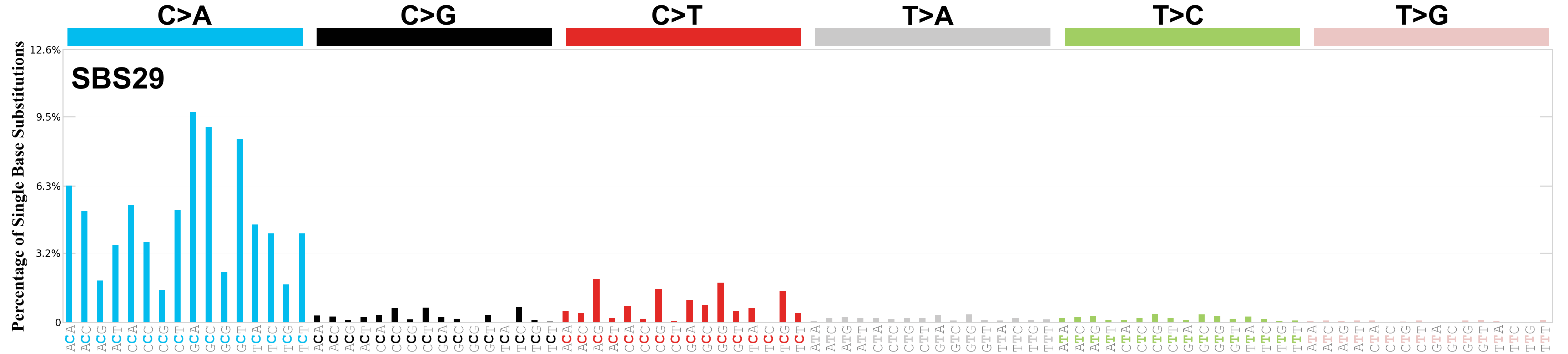
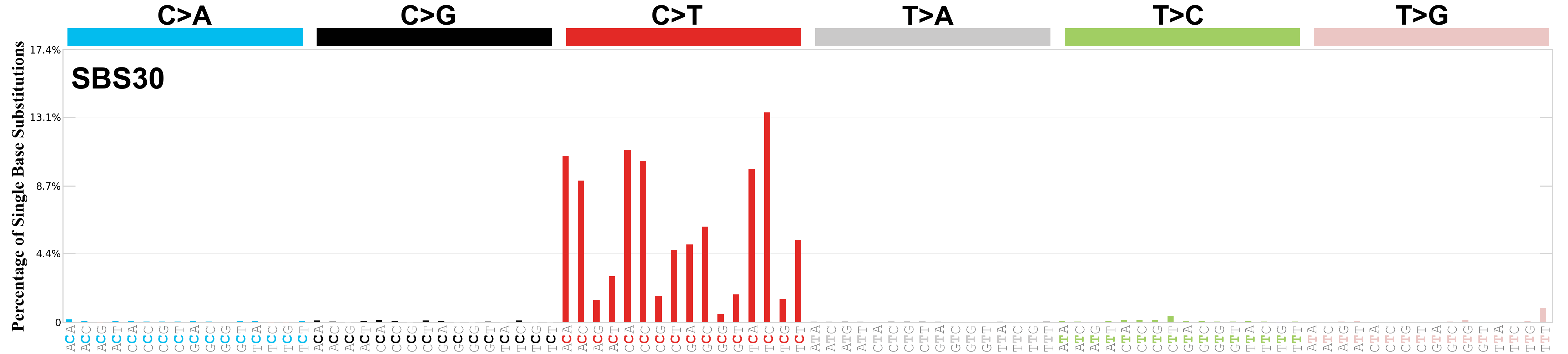
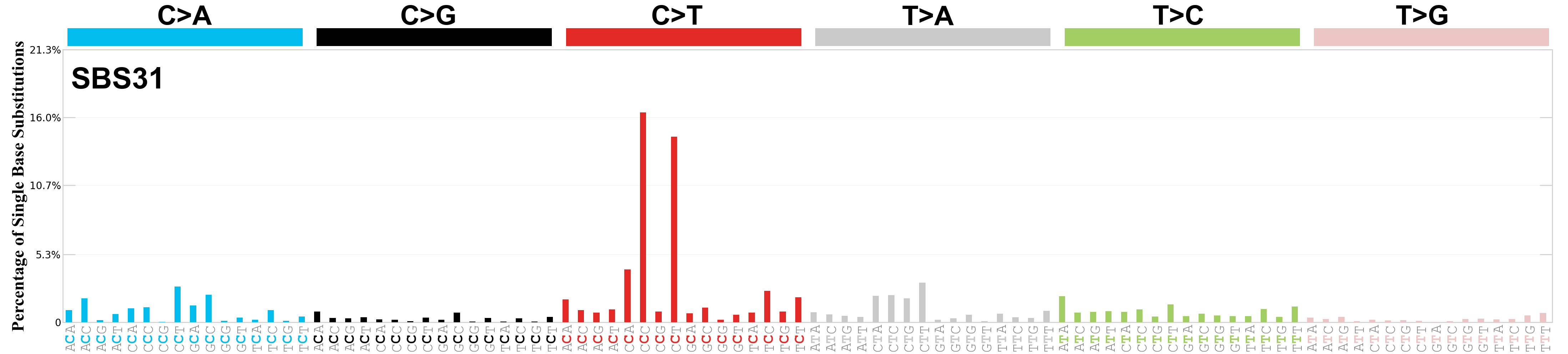
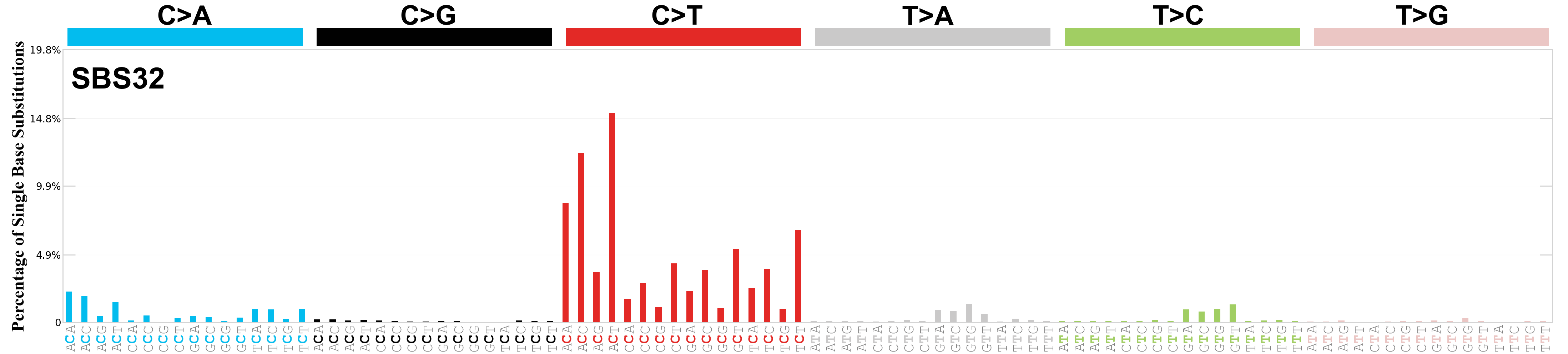
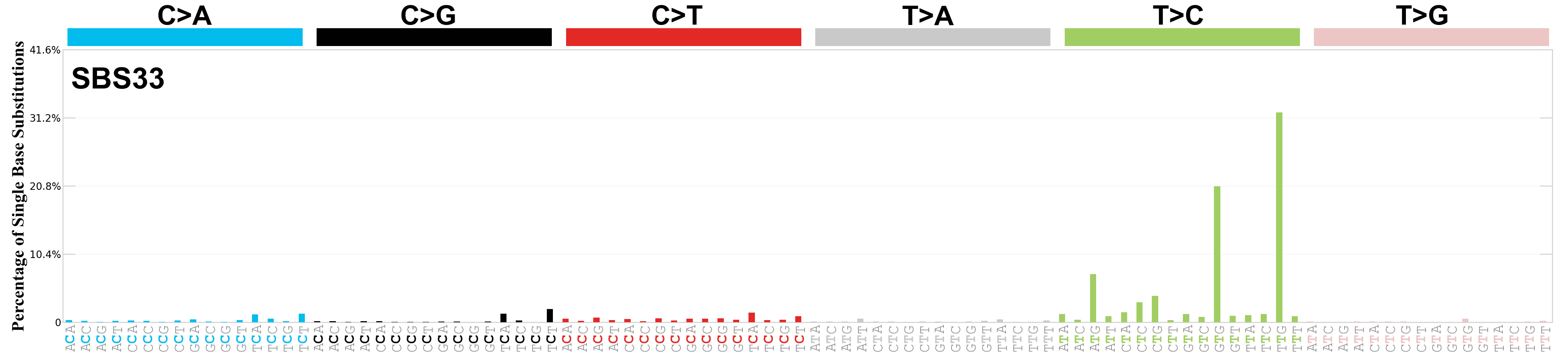
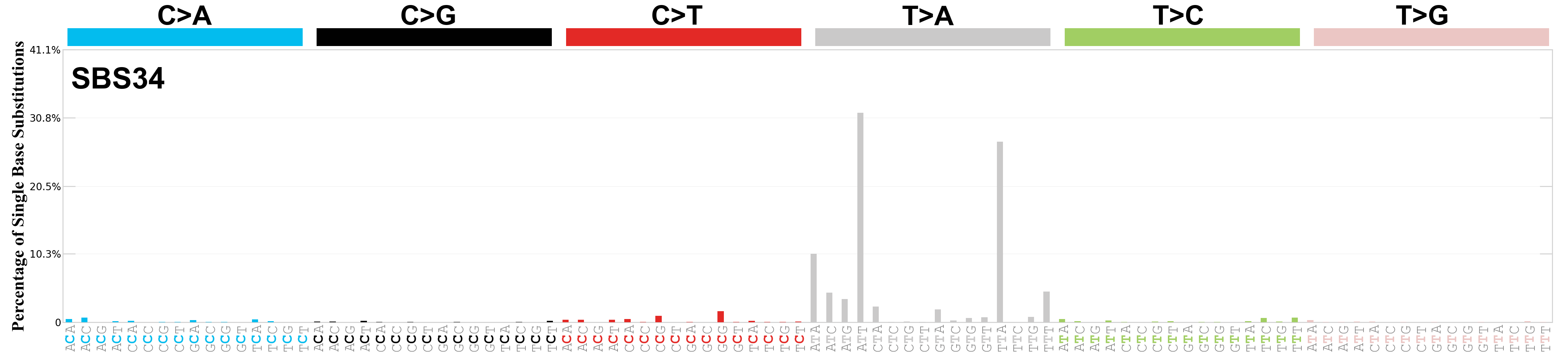
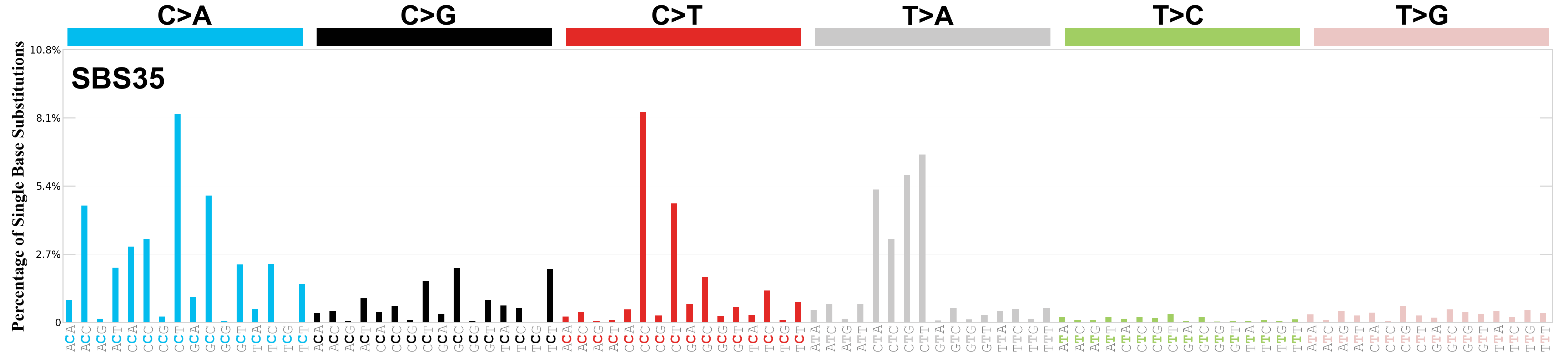
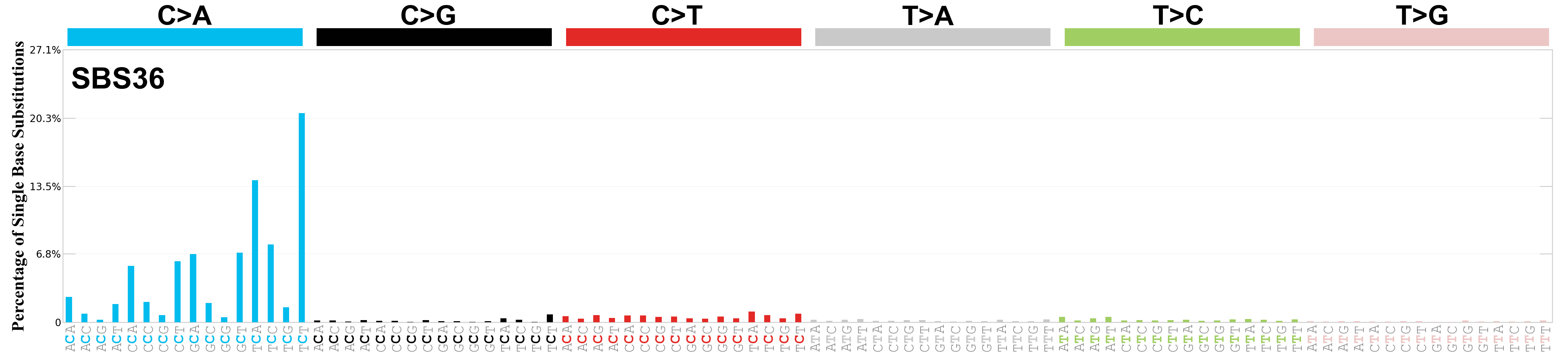
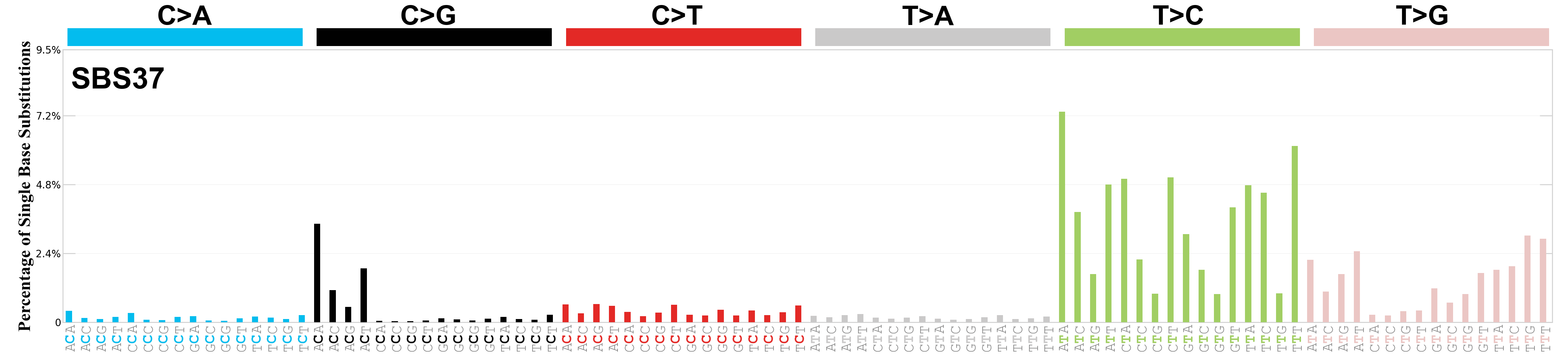
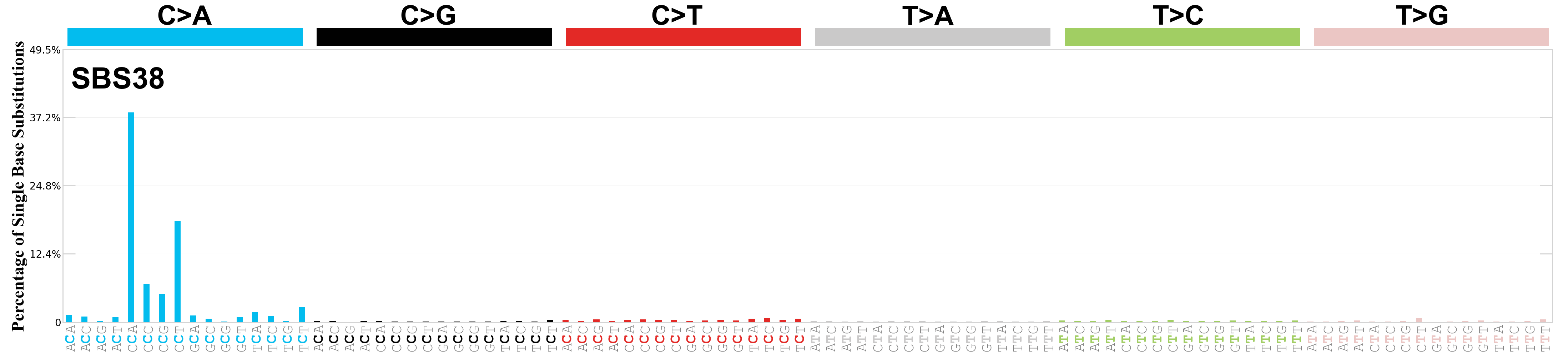
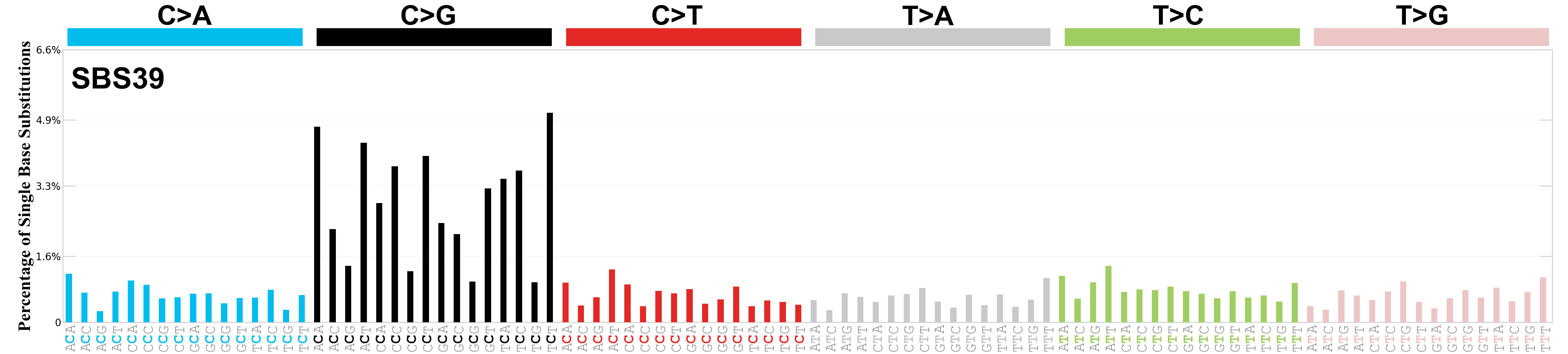
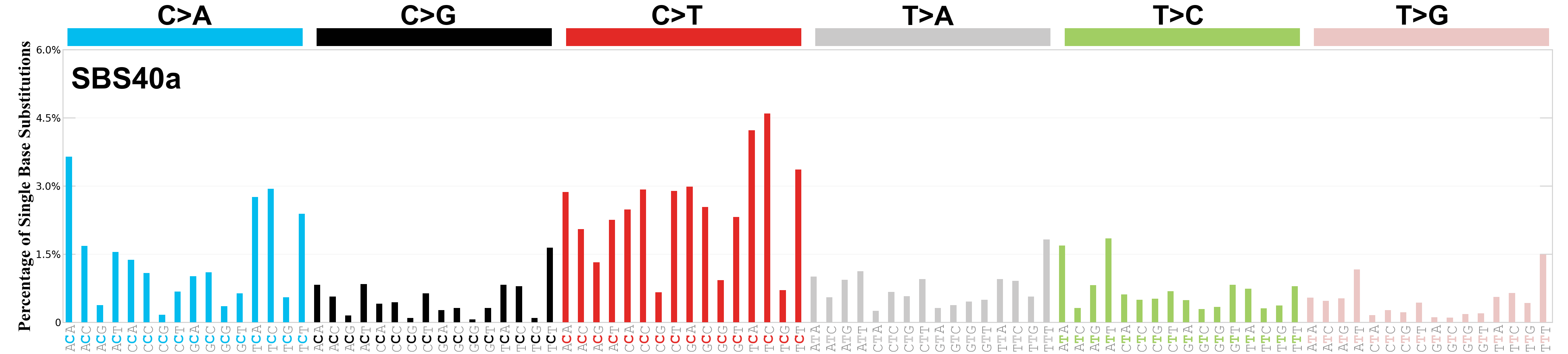
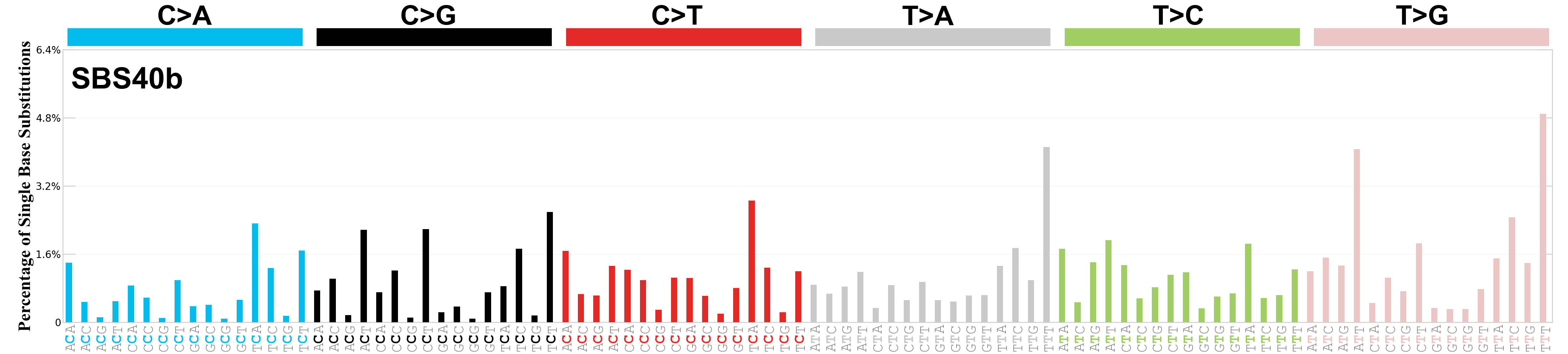
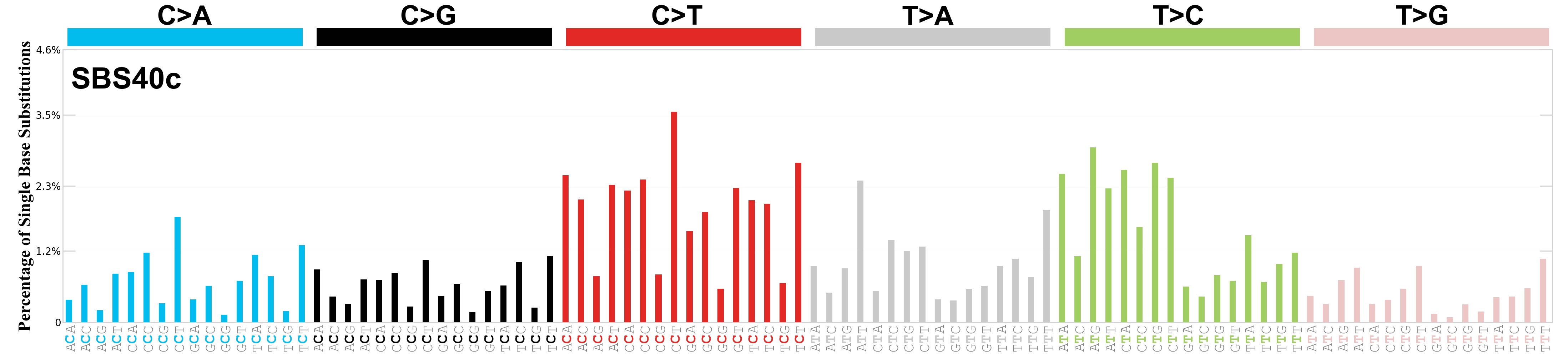
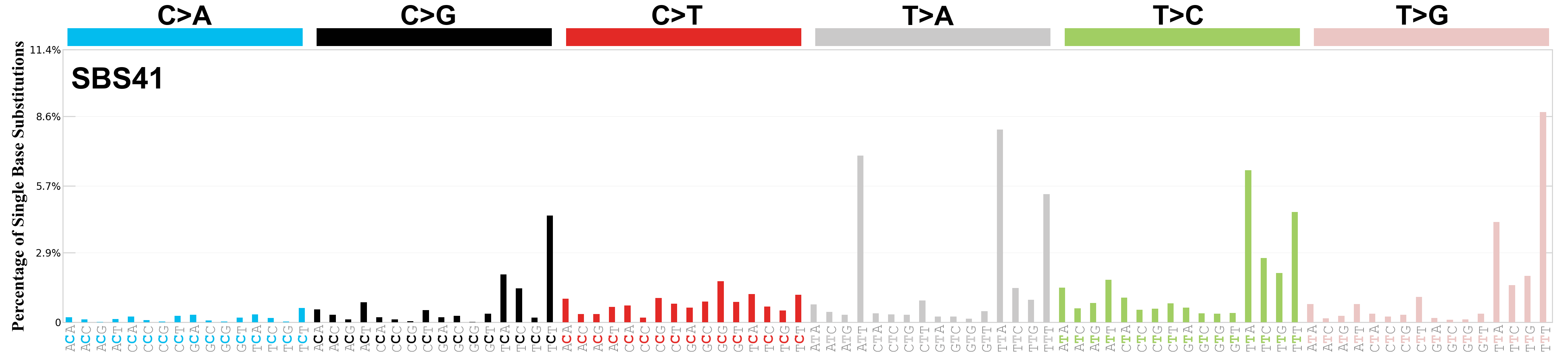
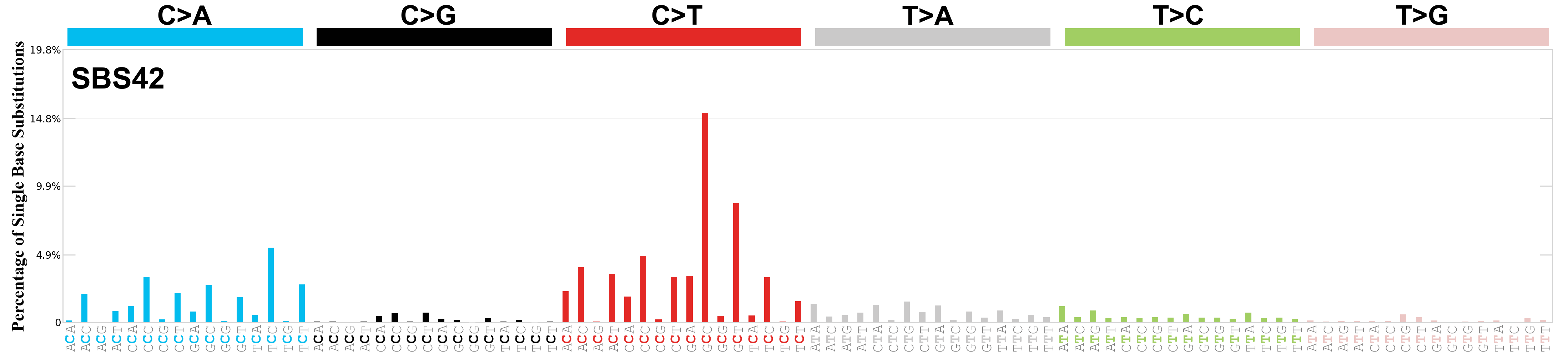
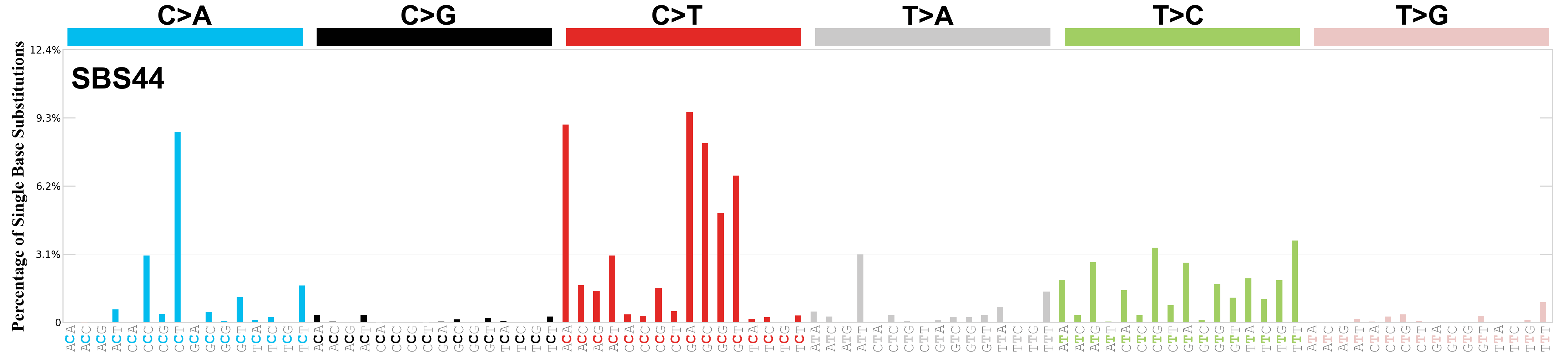
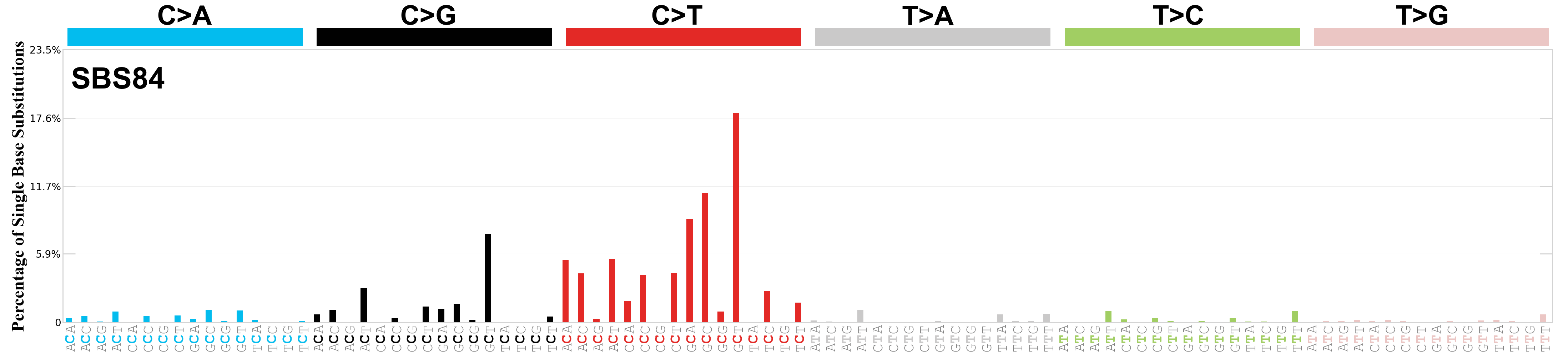
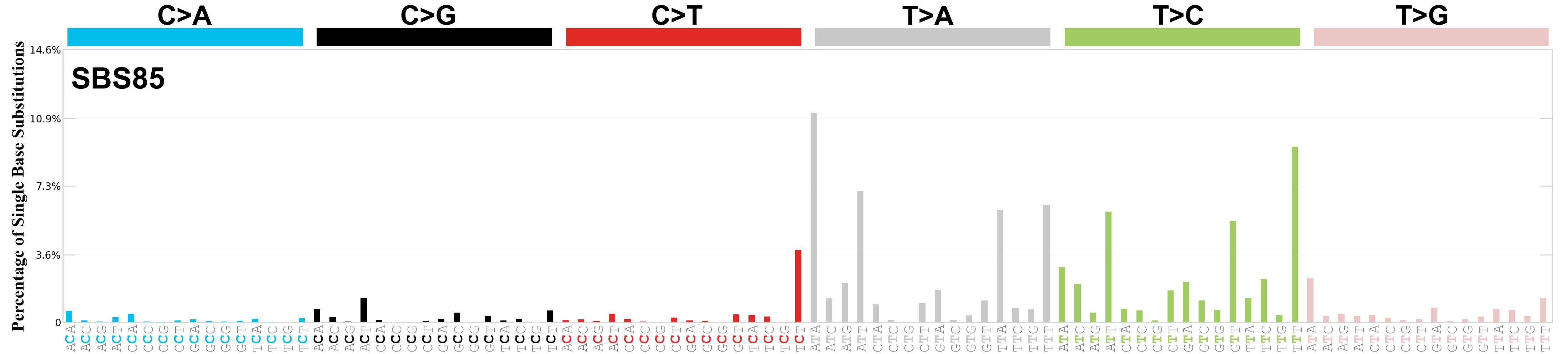
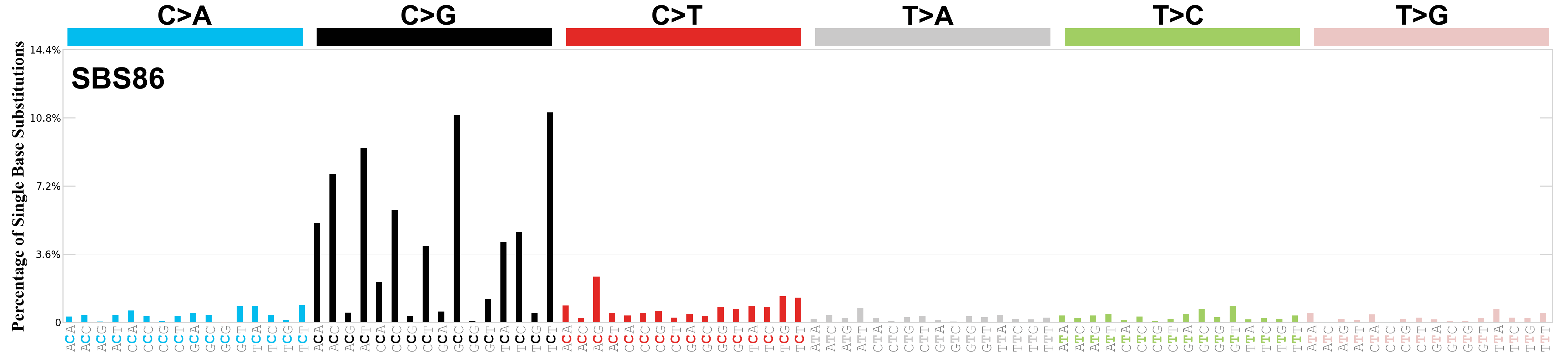
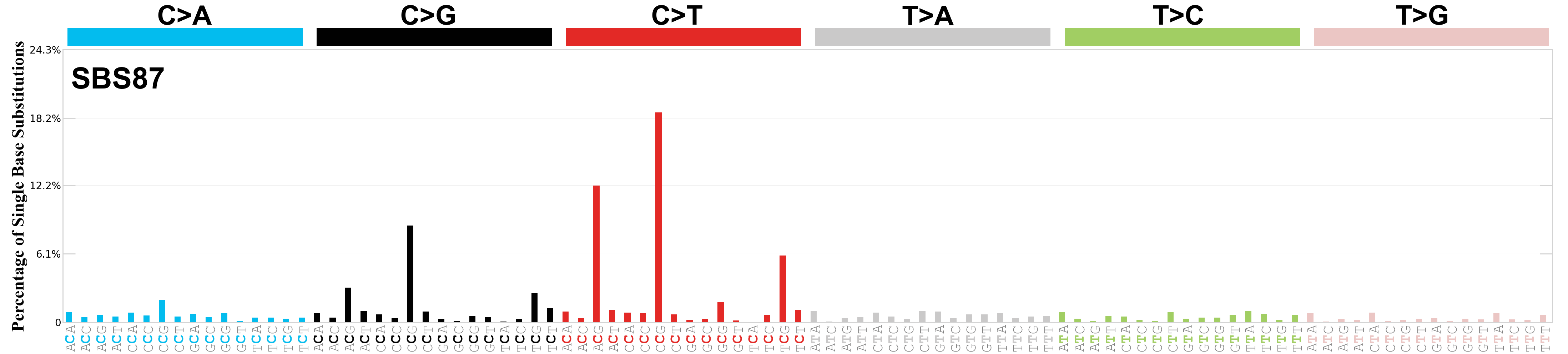
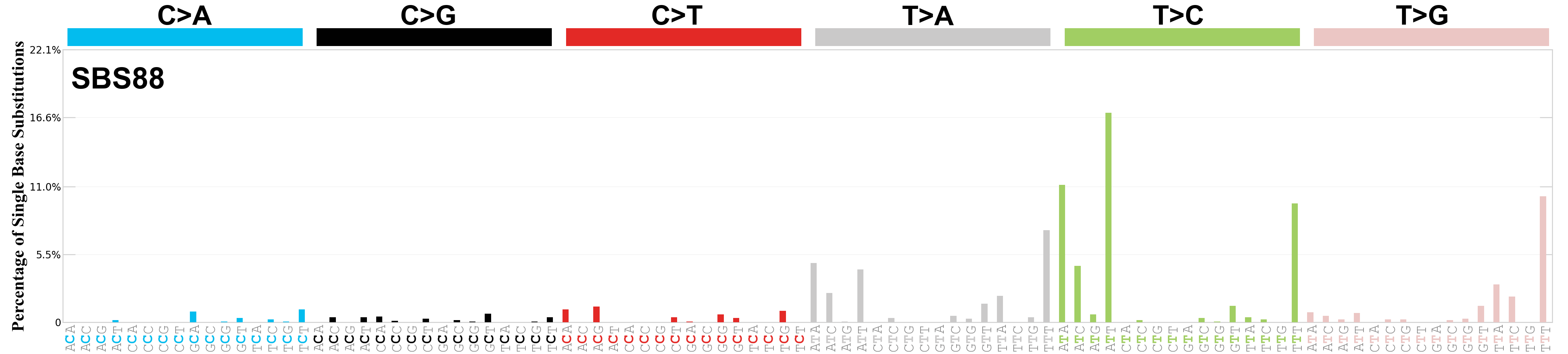
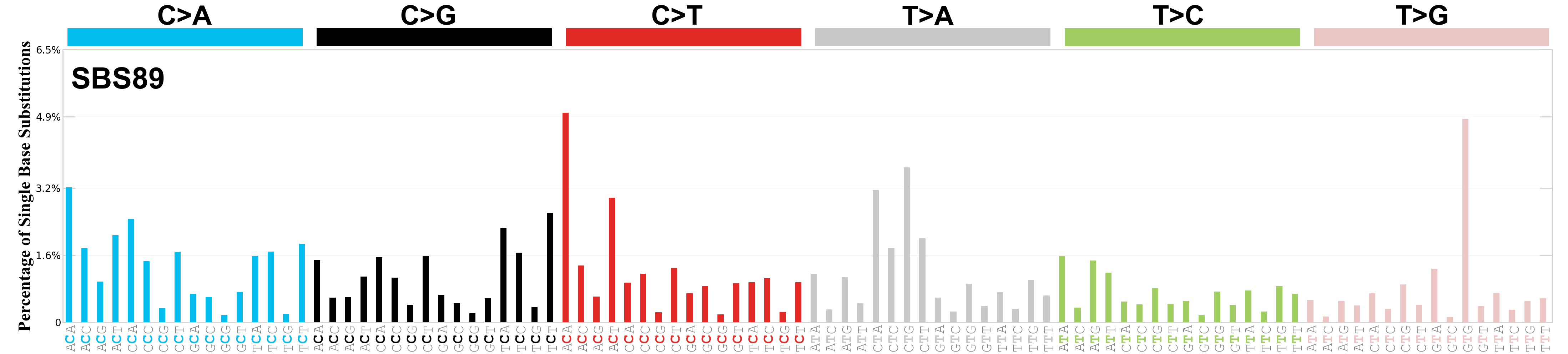
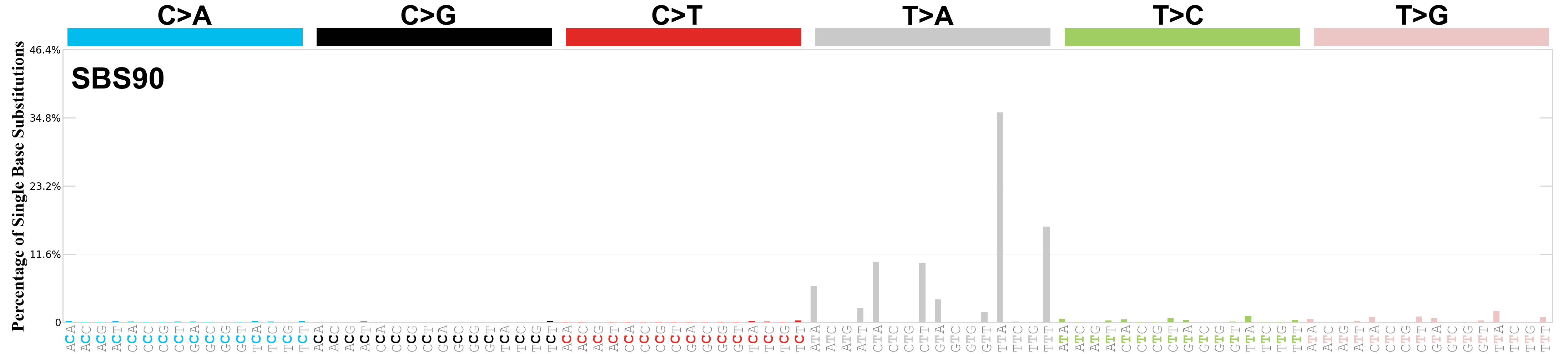
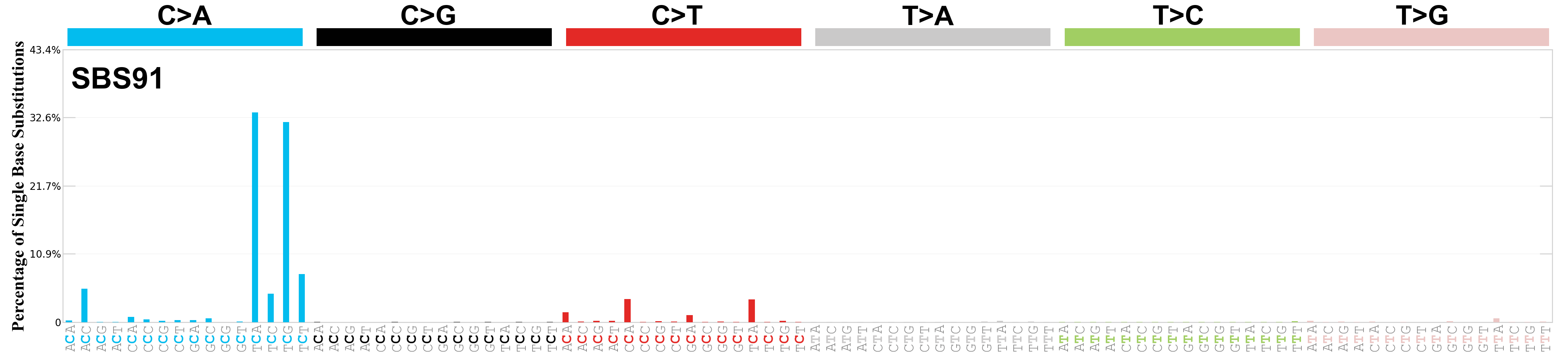
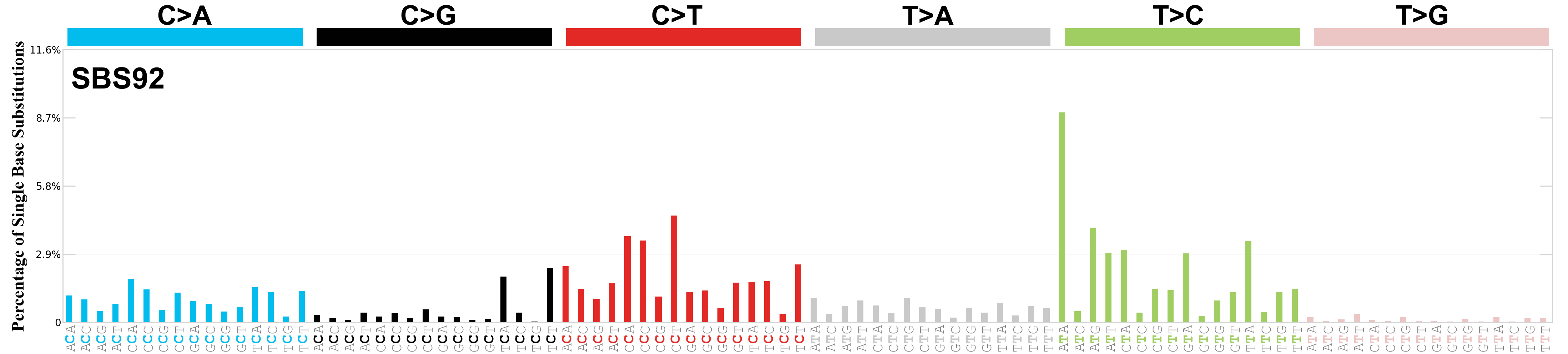
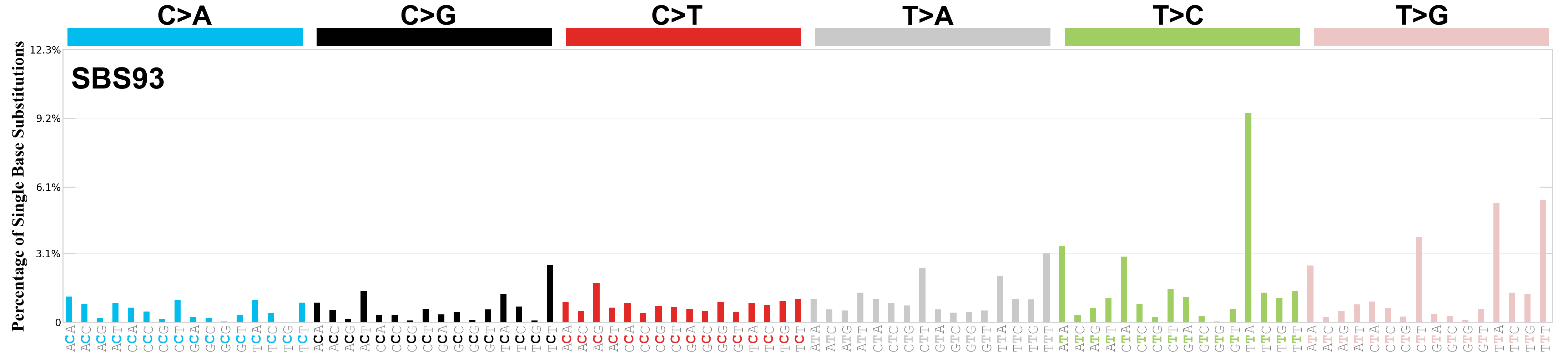
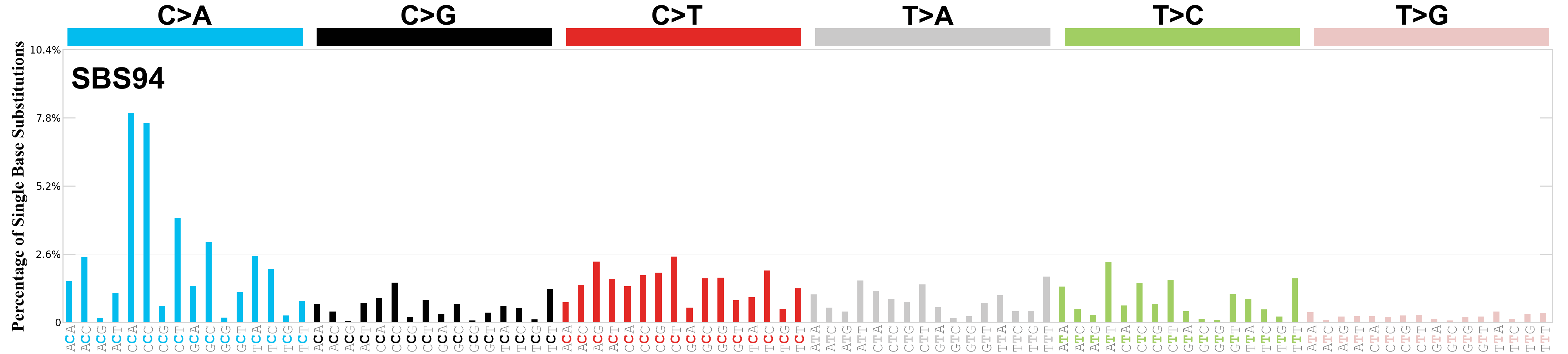
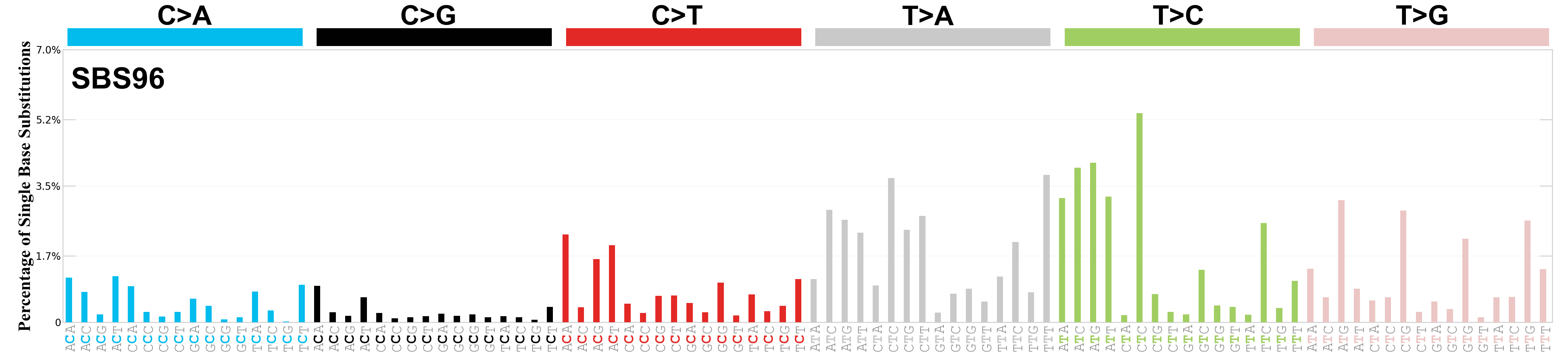
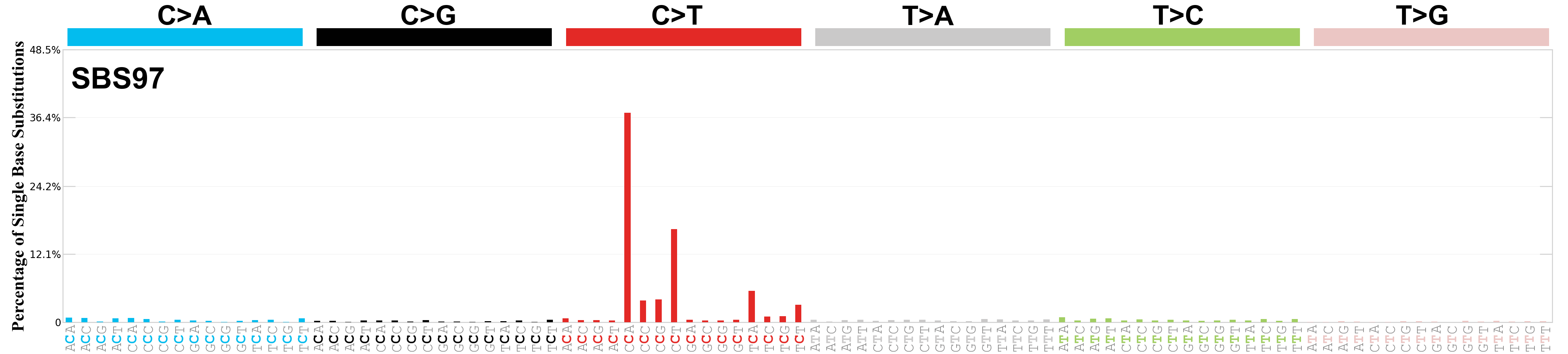
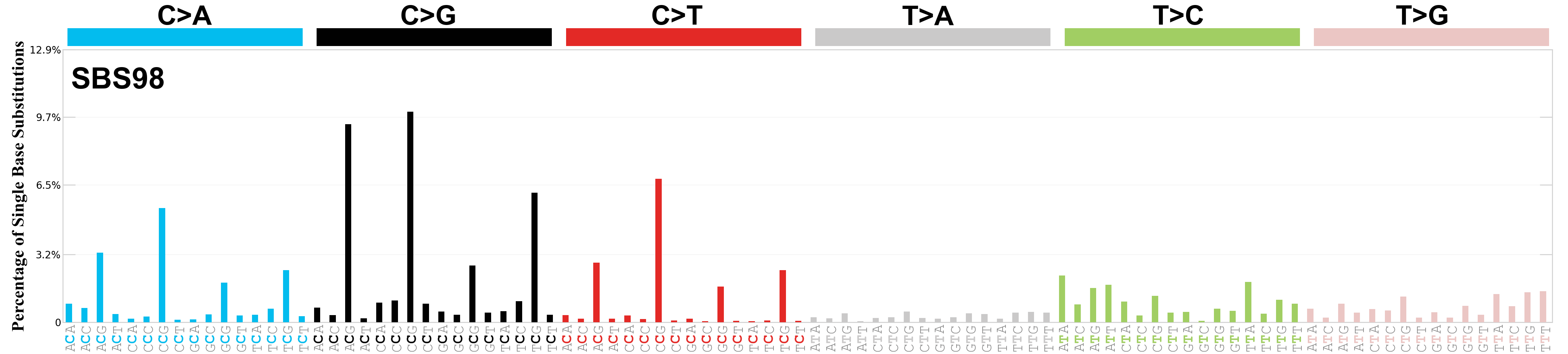
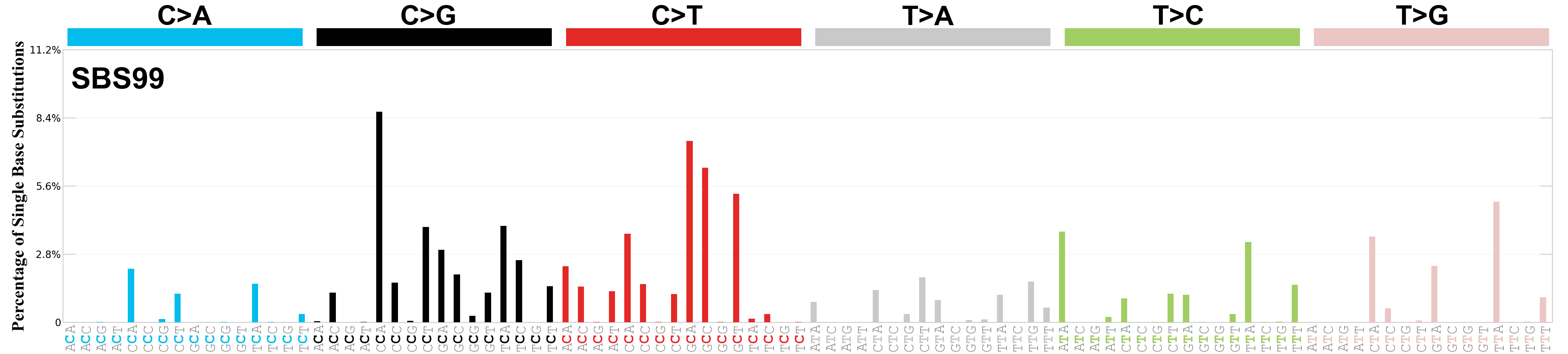
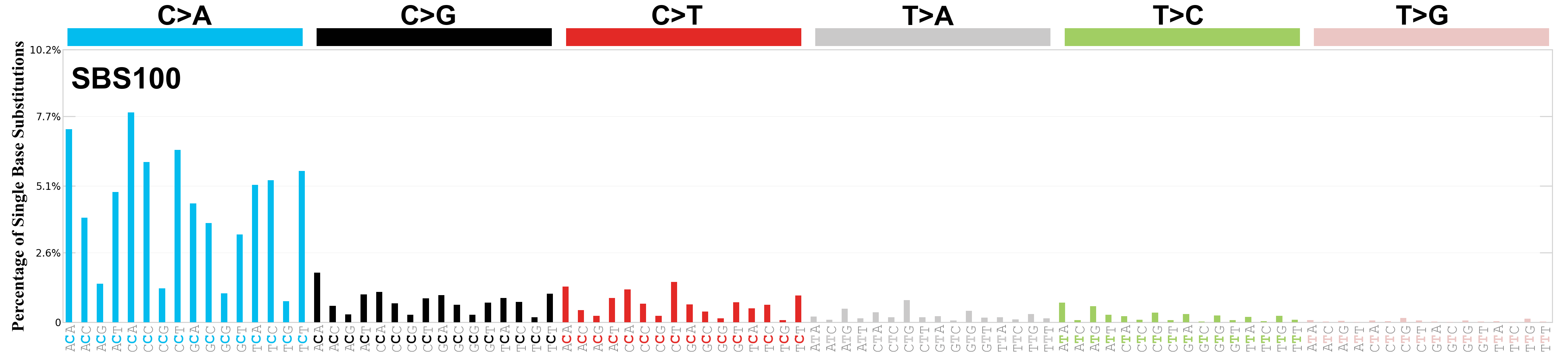
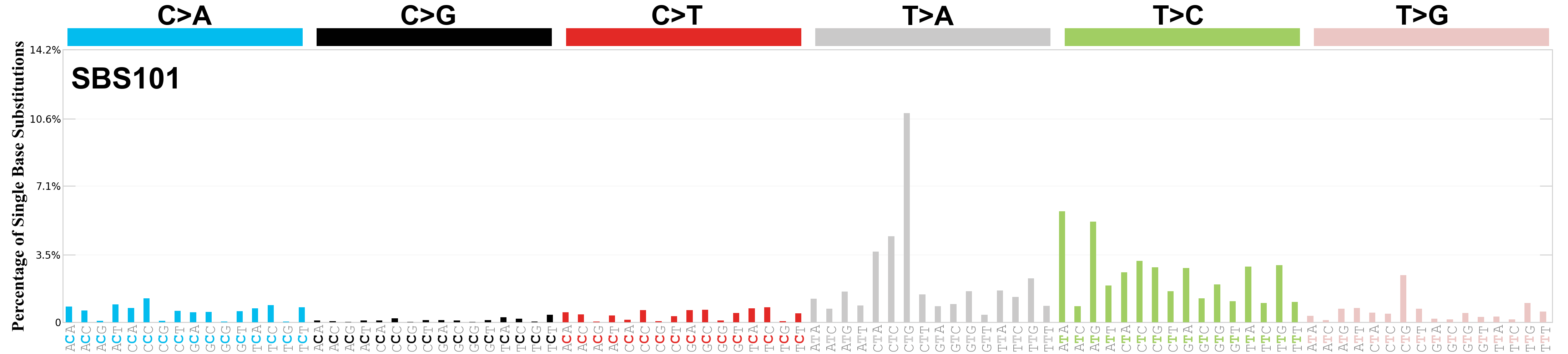
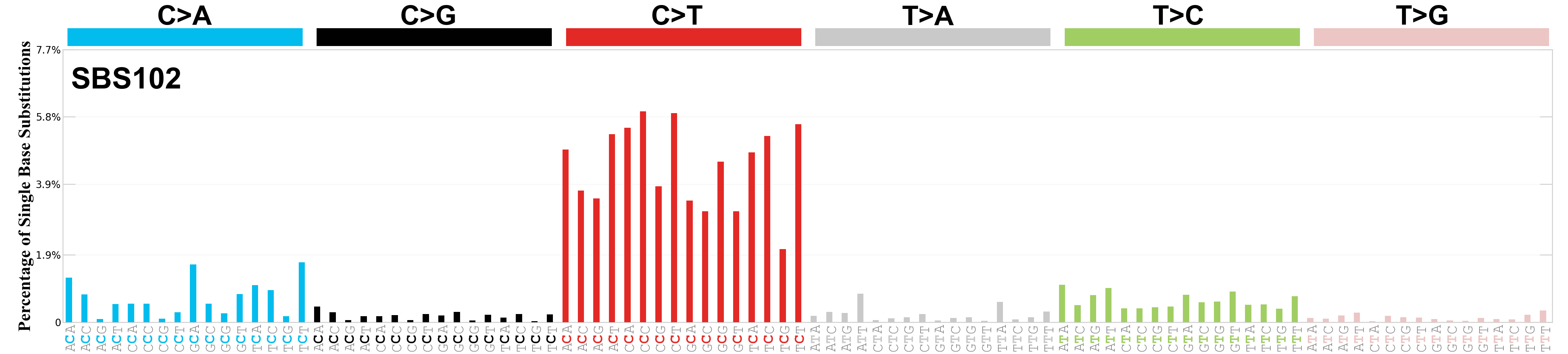
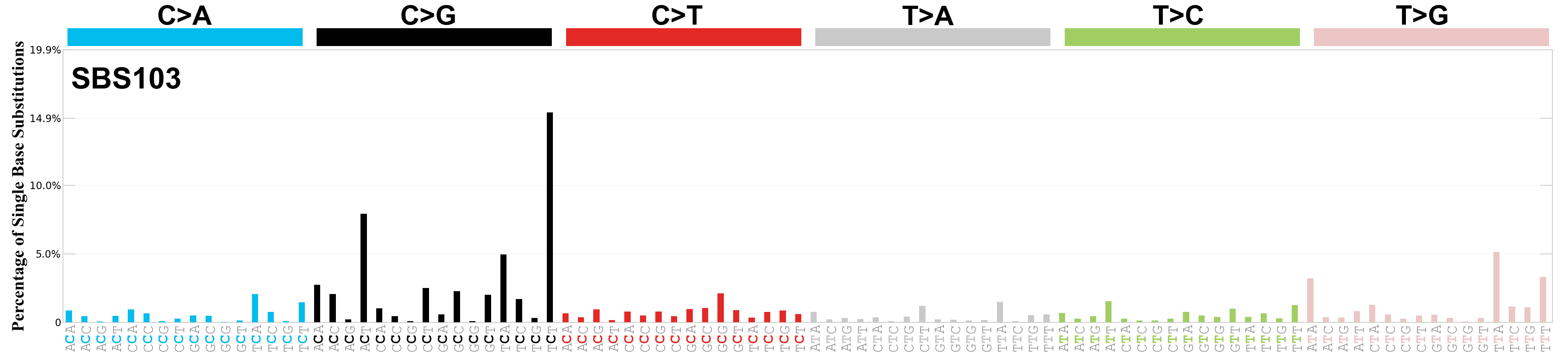
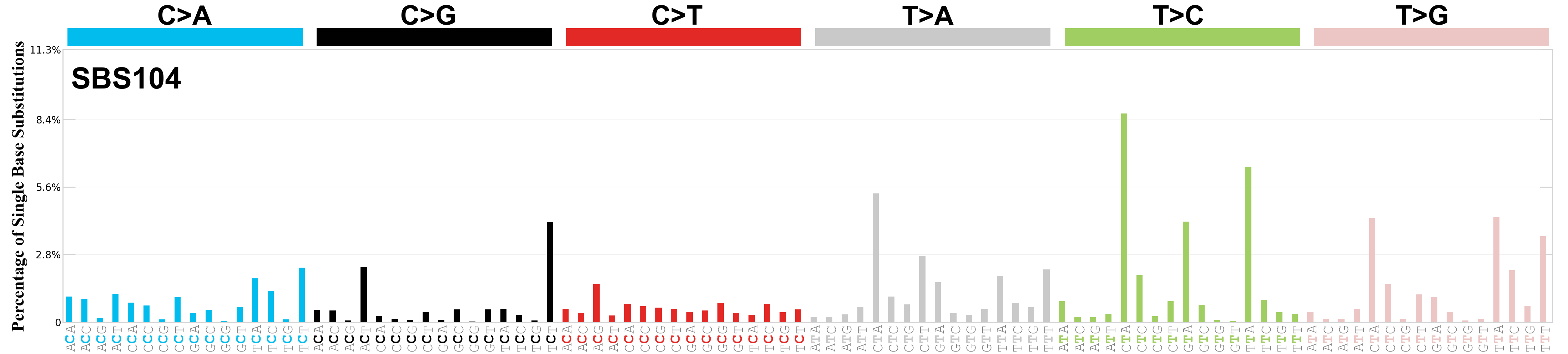
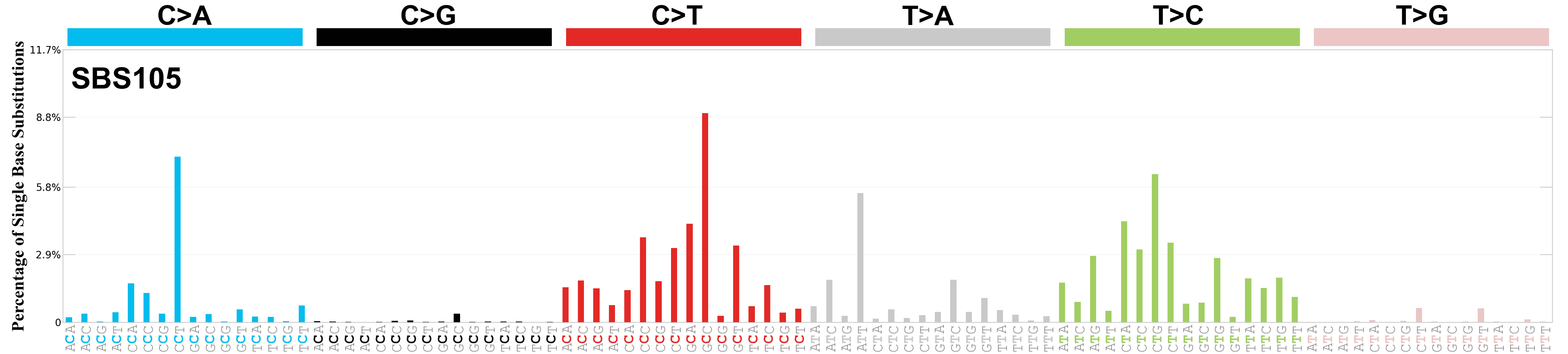
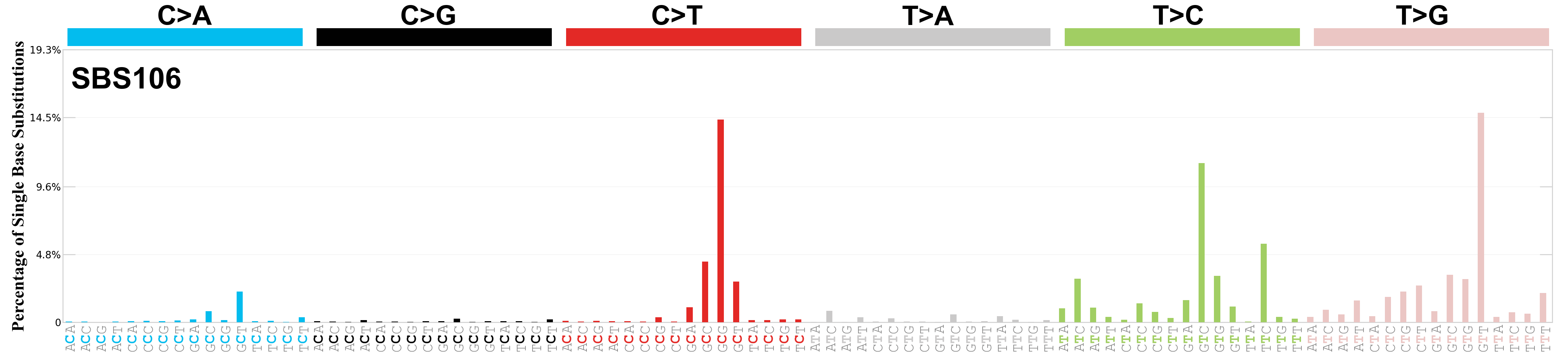
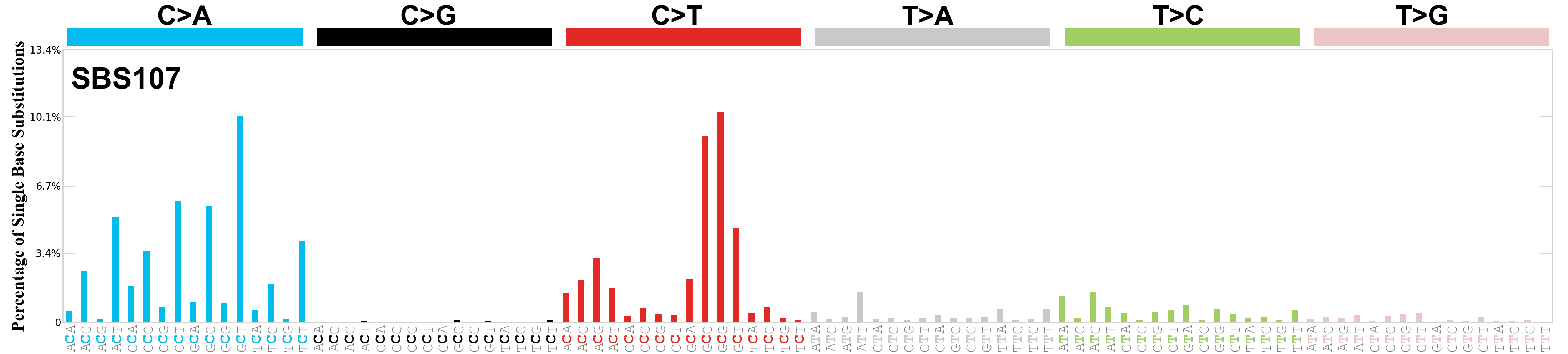
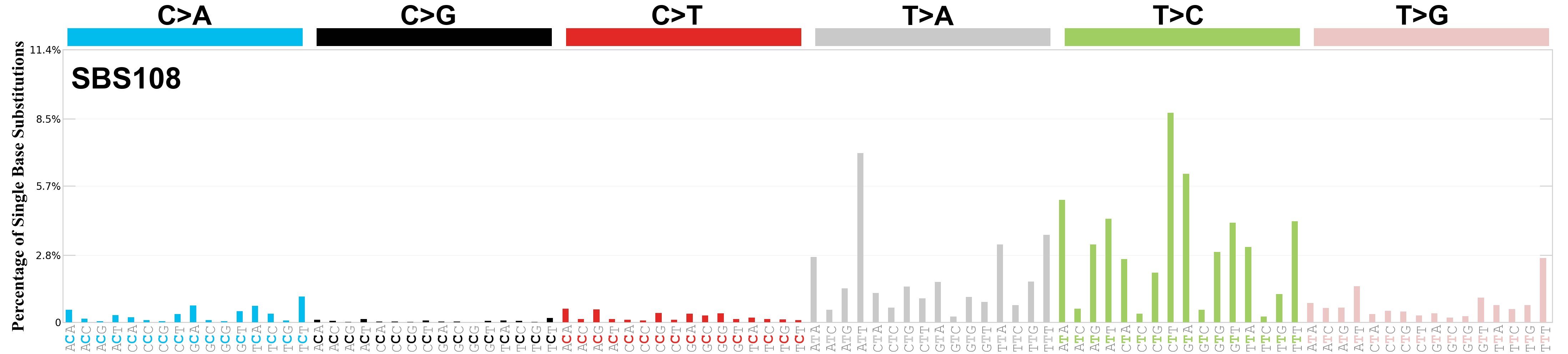
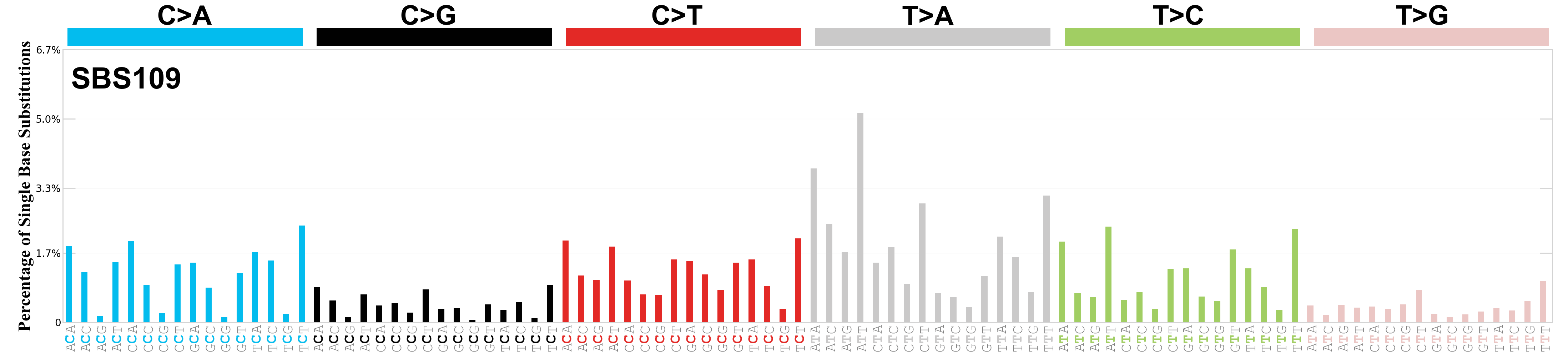
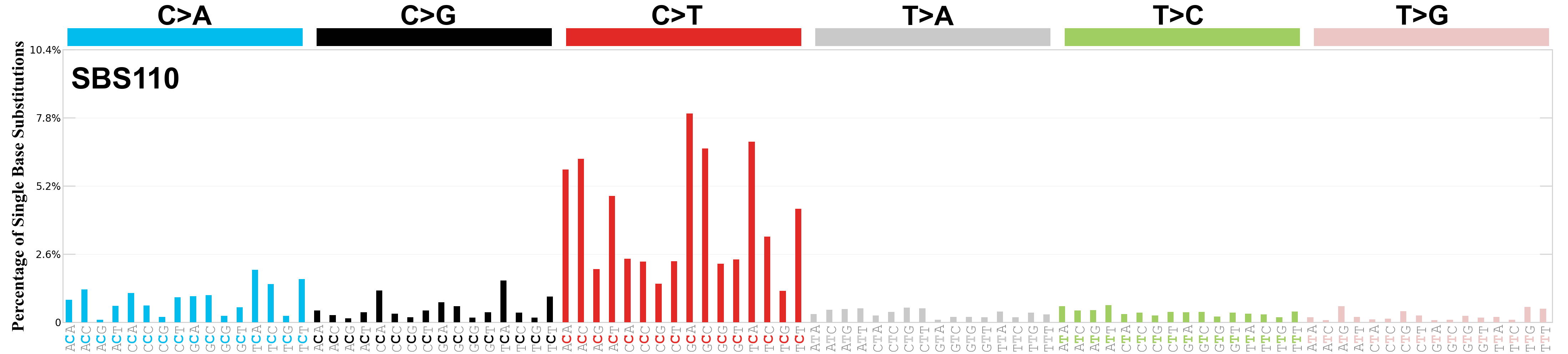
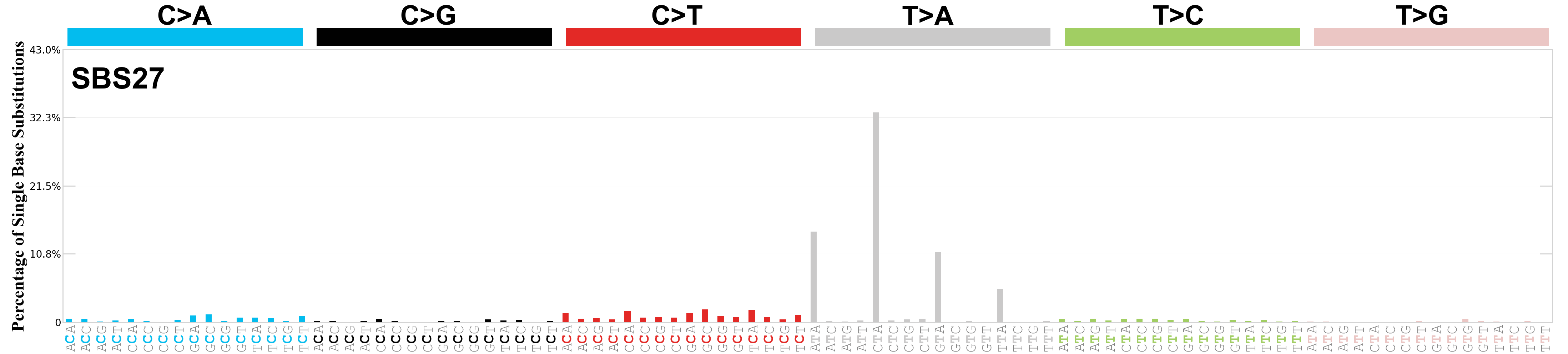
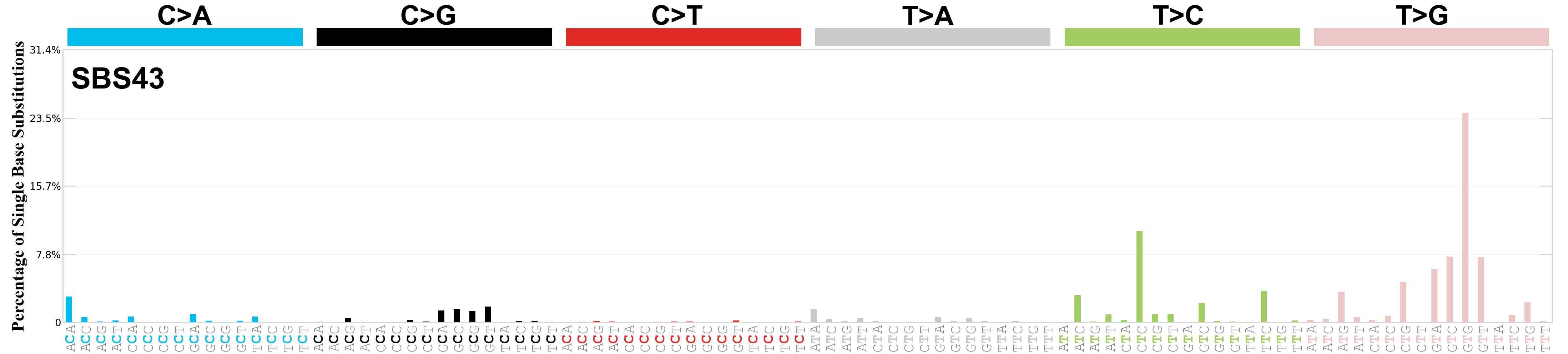
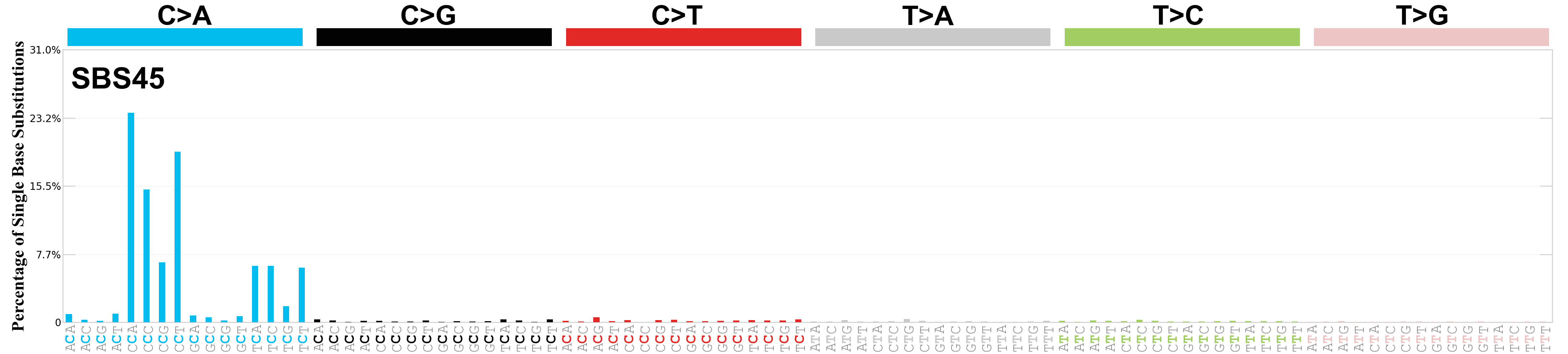
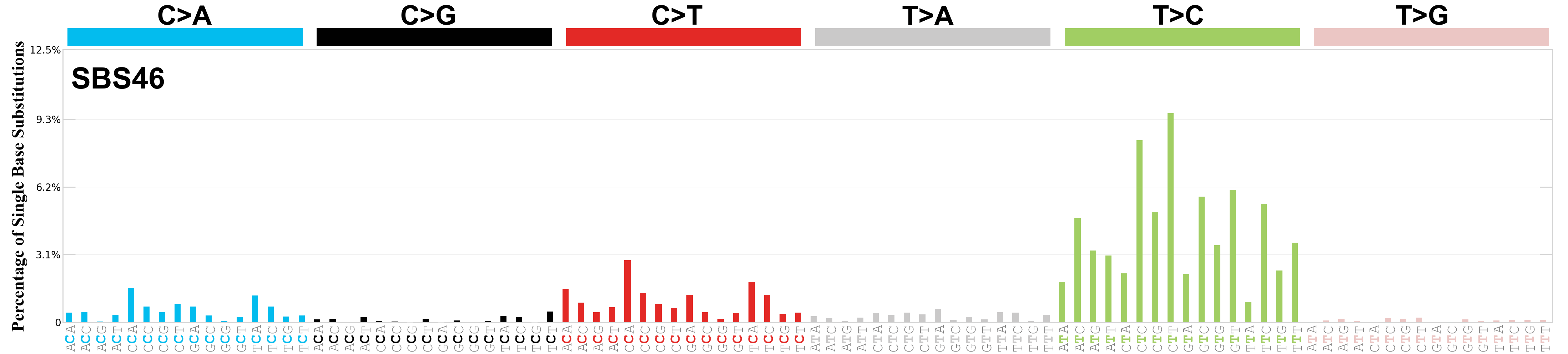
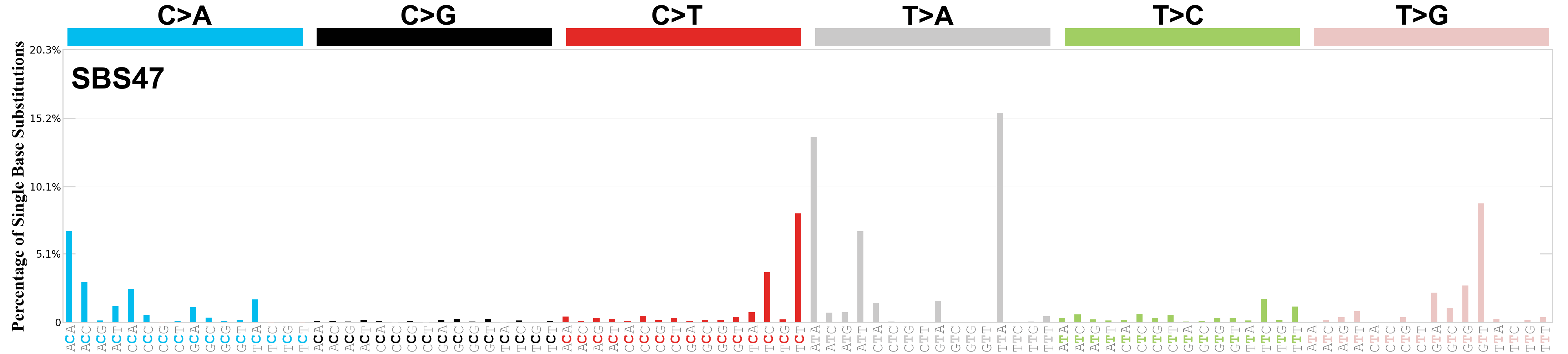
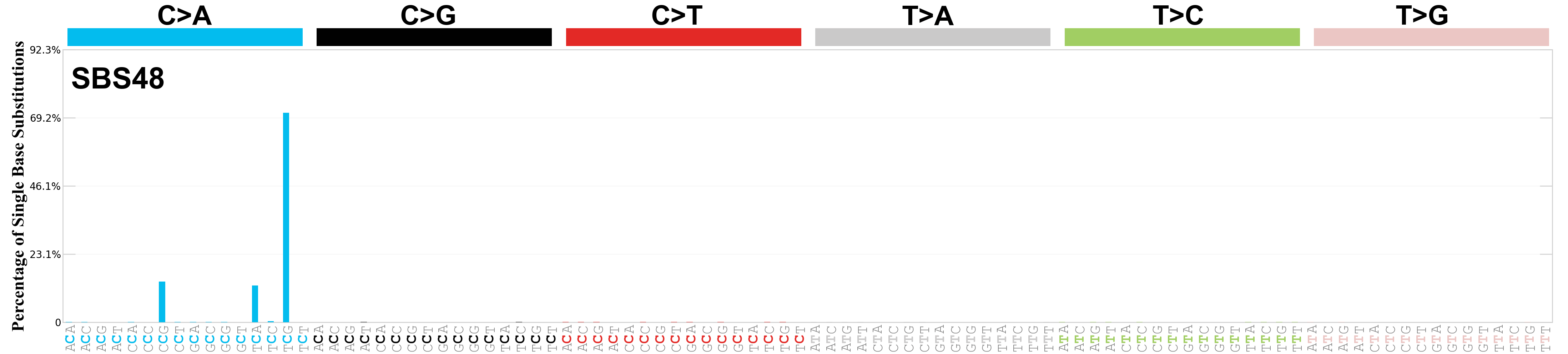
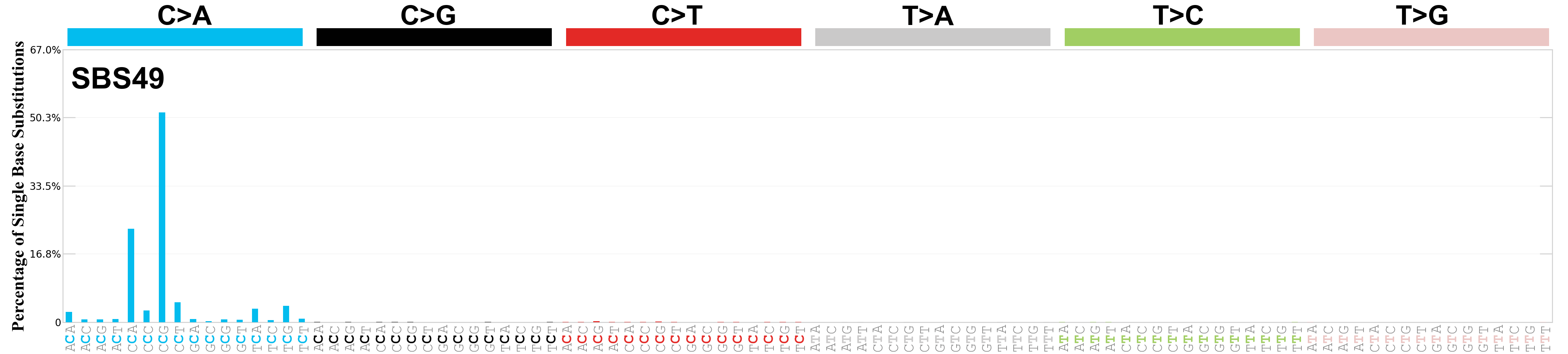
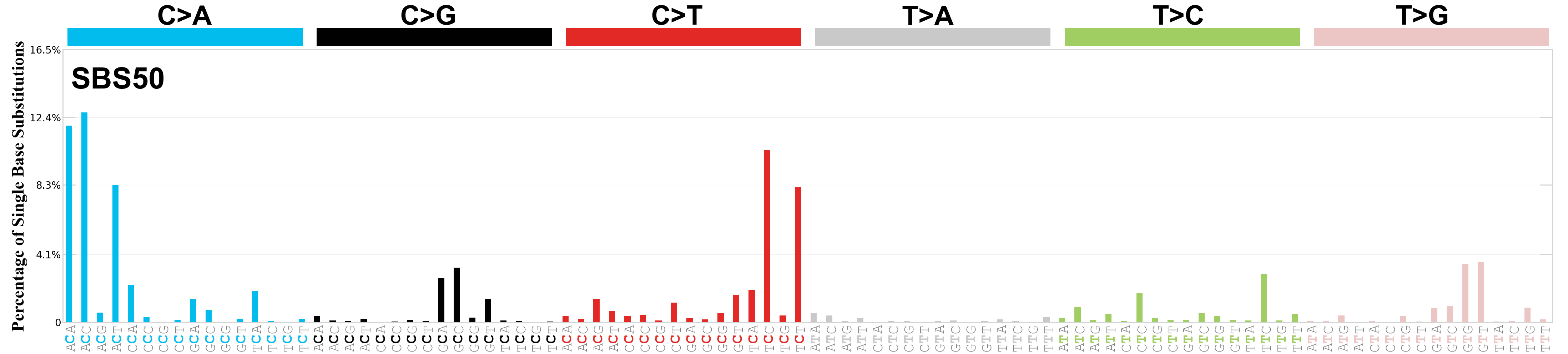
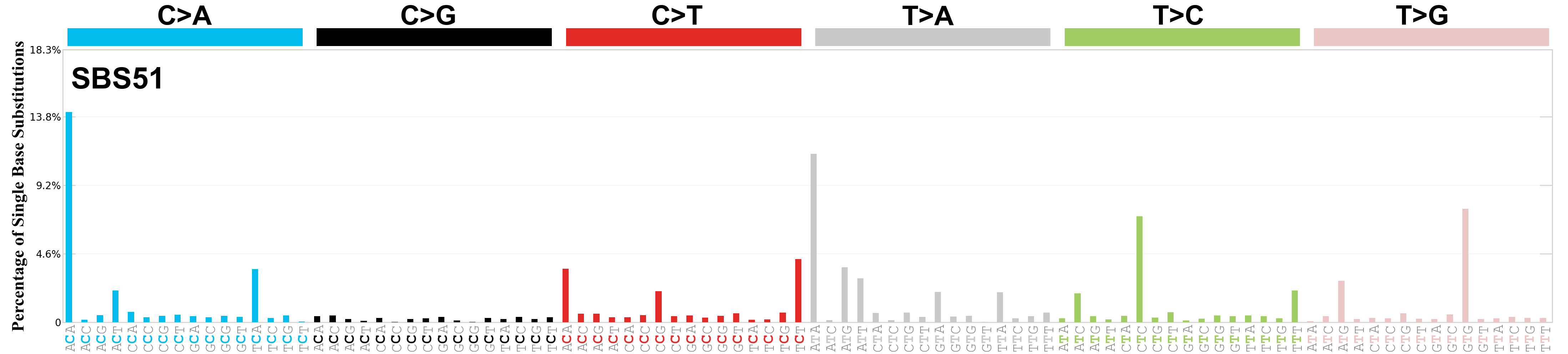
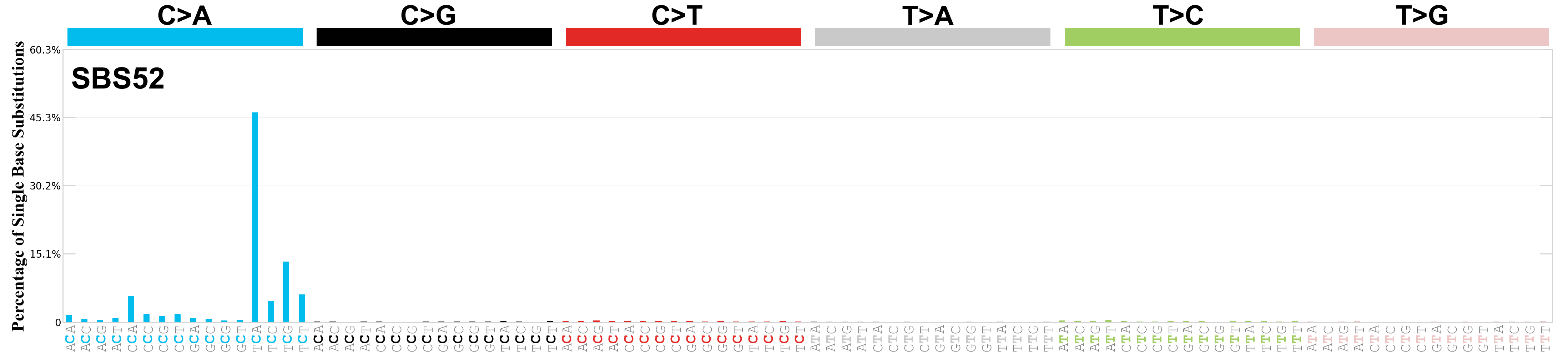
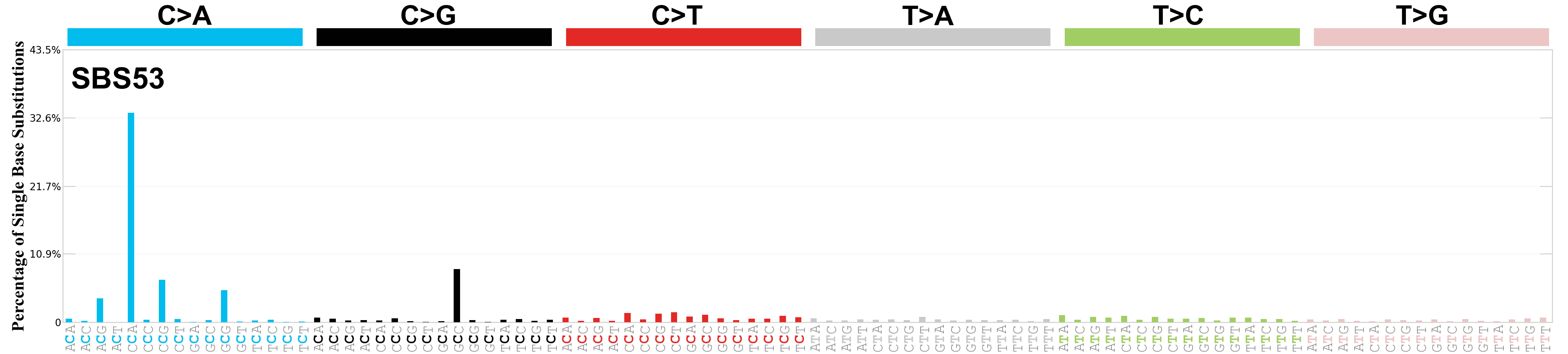
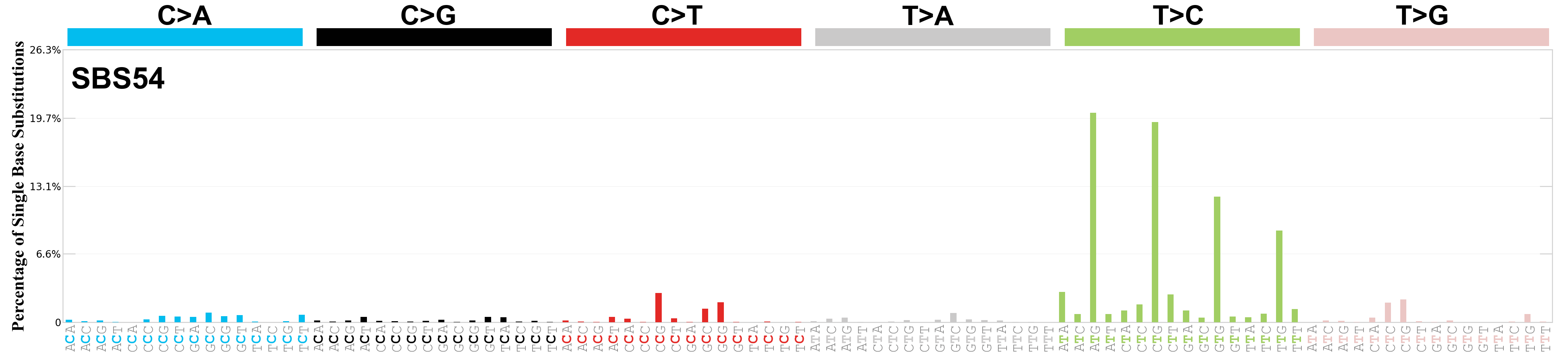
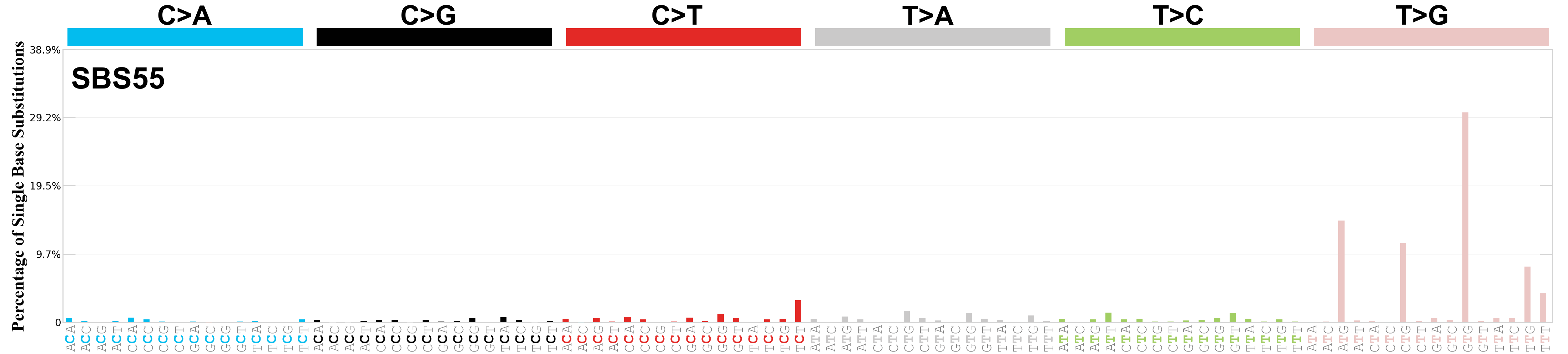
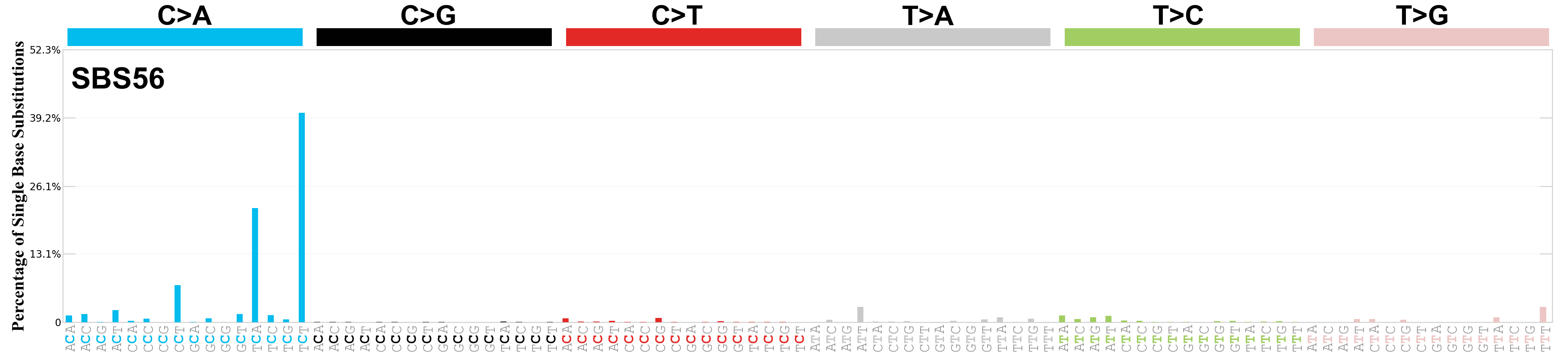
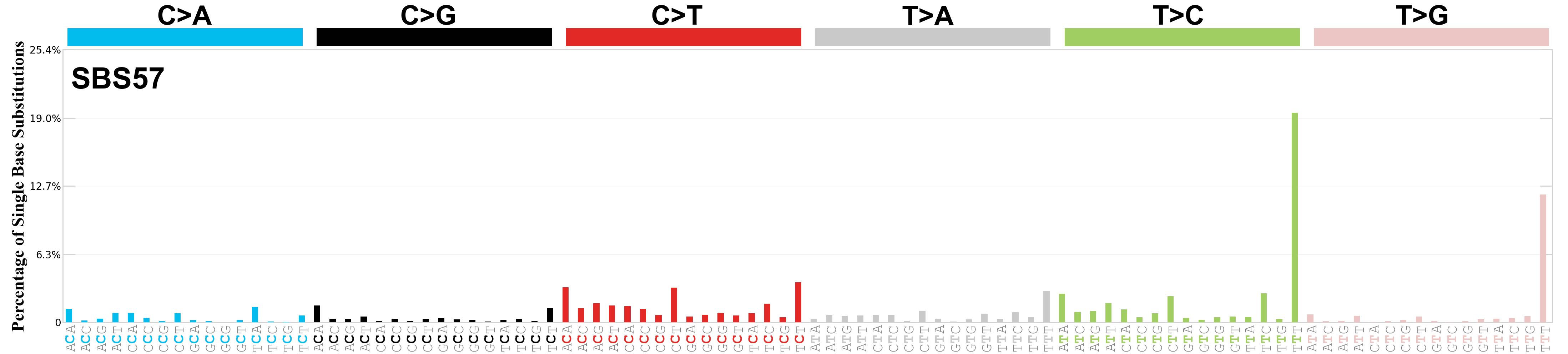
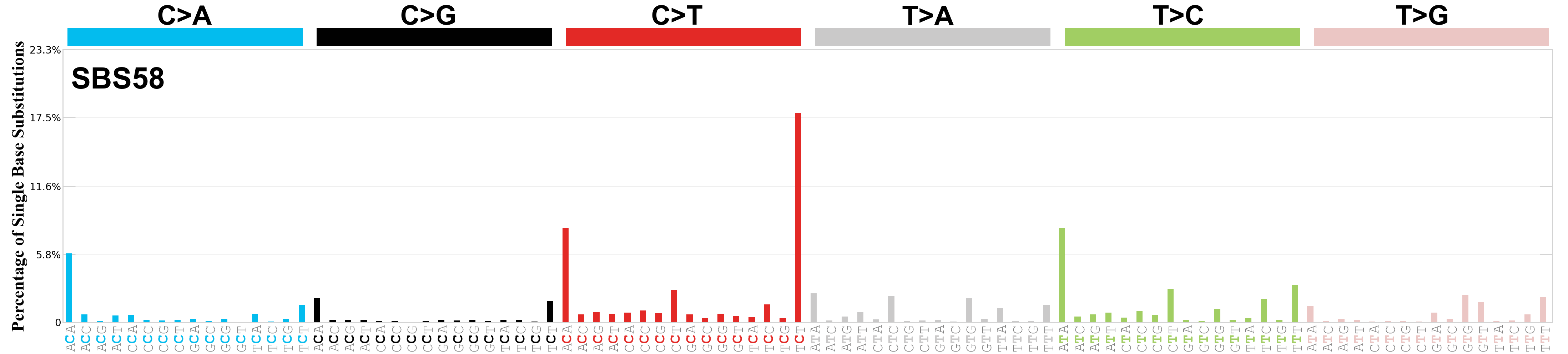
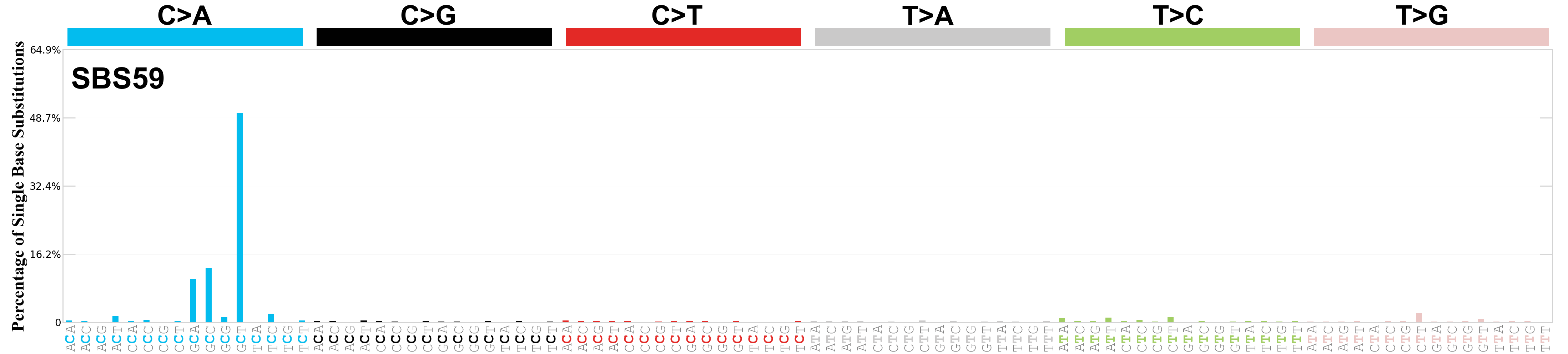
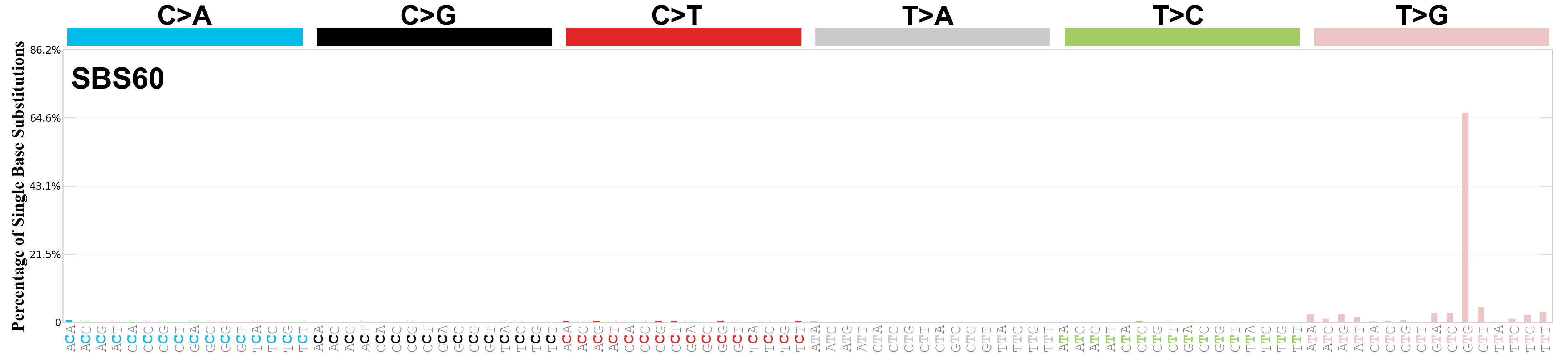
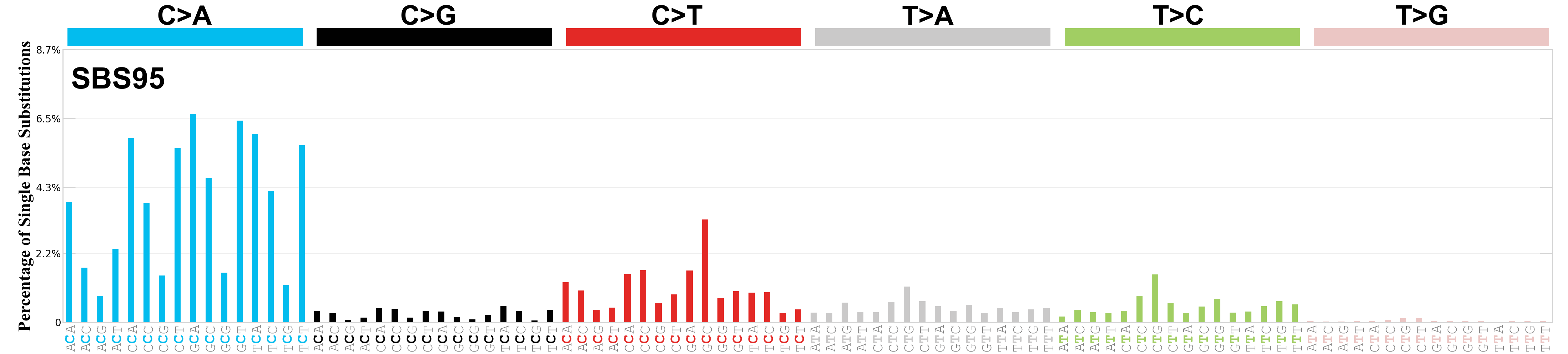
Single base substitutions (SBS), also known as single nucleotide variants, are defined as a replacement of a certain nucleotide base. Considering the pyrimidines of the Watson-Crick base pairs, there are only six different possible substitutions: C>A, C>G, C>T, T>A, T>C, and T>G. These SBS classes can be further expanded considering the nucleotide context.
Current SBS signatures have been identified using 96 different contexts, considering not only the mutated base, but also the bases immediately 5’ and 3’.
Click on any signature below to learn more about its details.
Methods
Signature extraction methods
With a few exceptions, the current set of reference signatures were extracted using SigProfiler (as described in Alexandrov, L.B. et al., 2020) from the 2,780 whole-genome variant calls produced by the ICGC/TCGA Pan Cancer Analysis of Whole Genomes (PCAWG) Network. The stability and reproducibility of the signatures were assessed on somatic mutations from an additional 1,865 whole genomes and 19,184 exomes. All input data and references for original sources are available from synapse.org ID syn11801889.
COSMIC mutational signatures are available in numerical form in our data downloads page.