Human Cancer Signatures
Version 3.6Copy Number Variations (CN) Signatures
About the collection
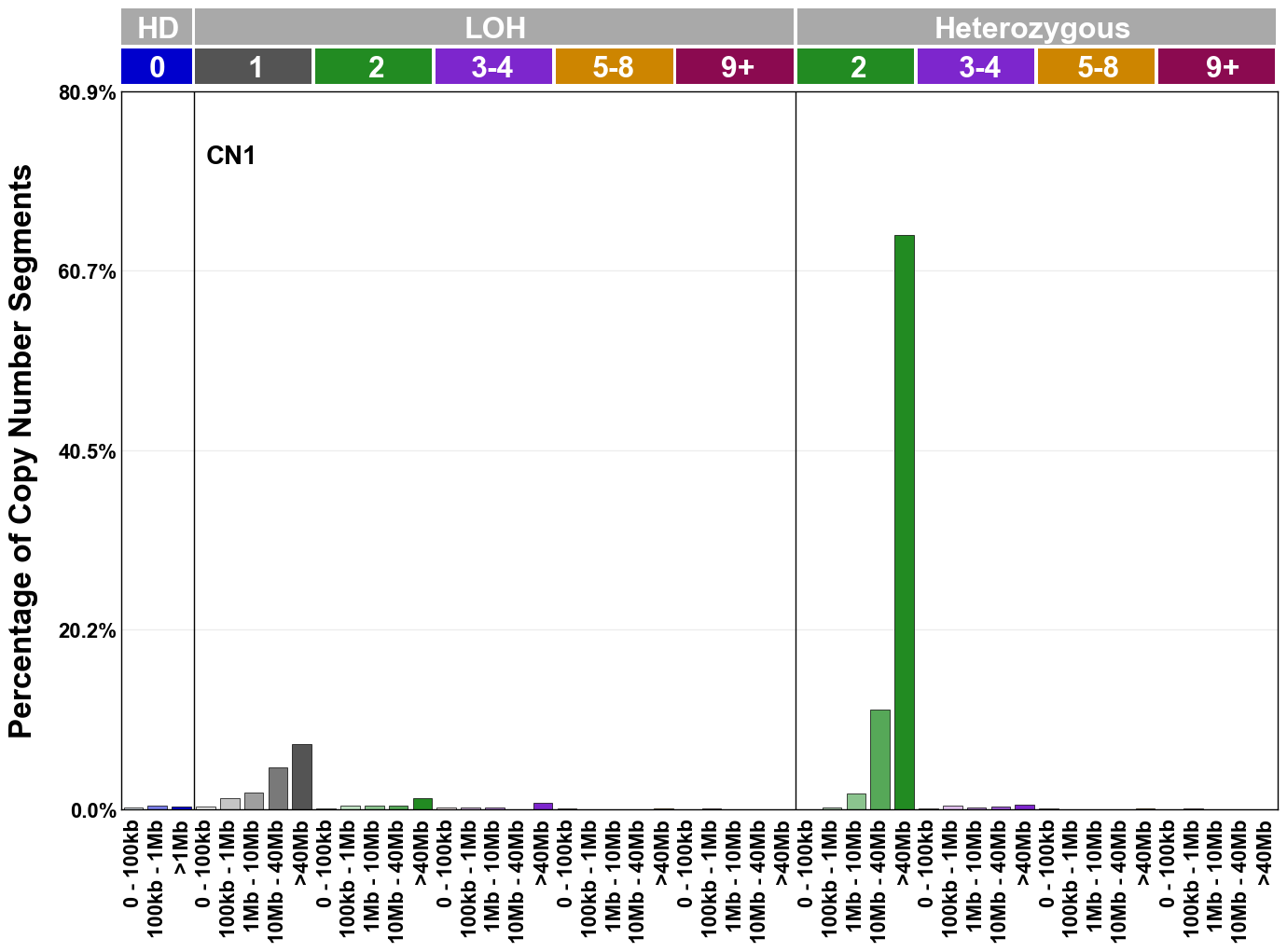
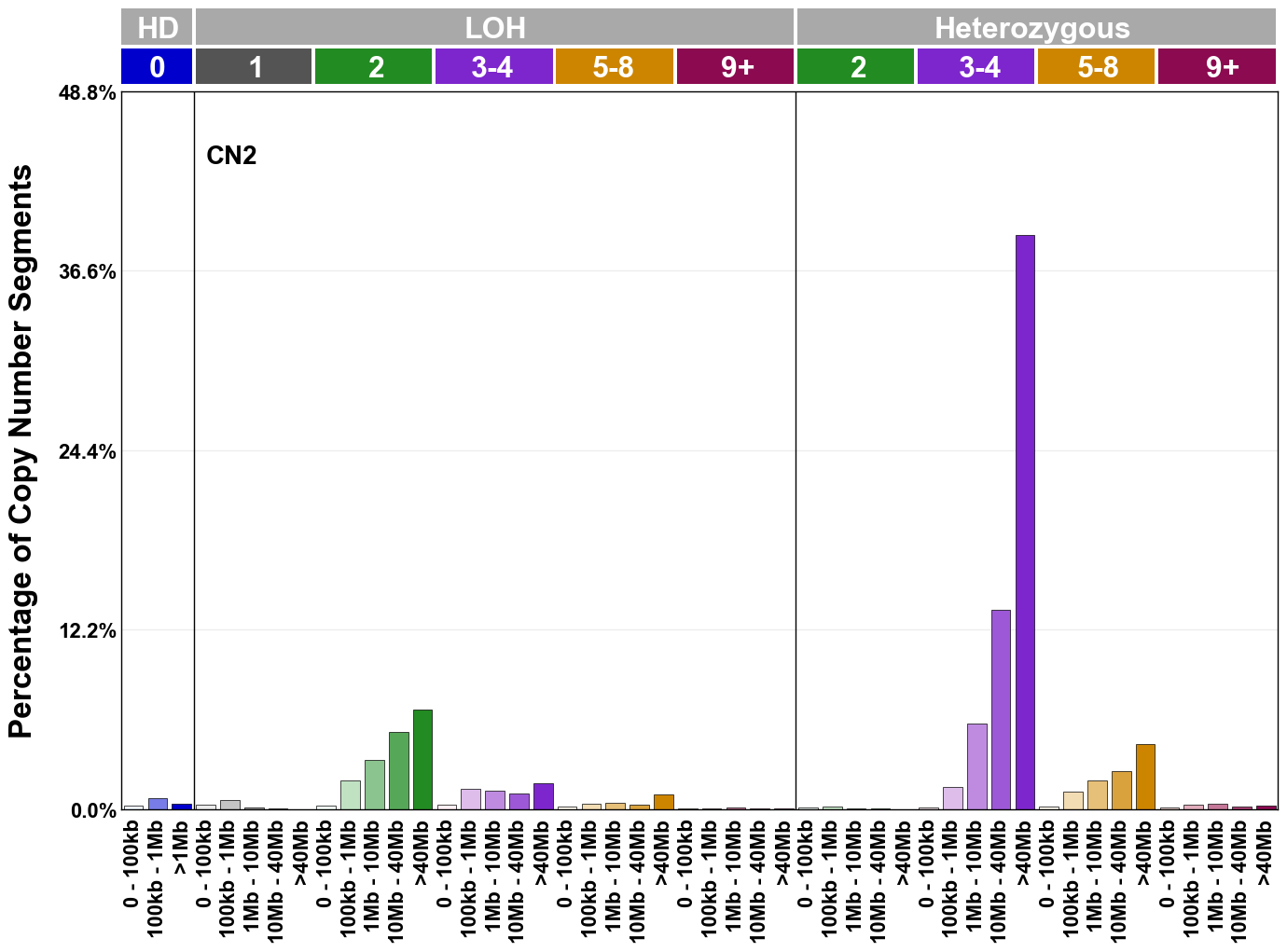
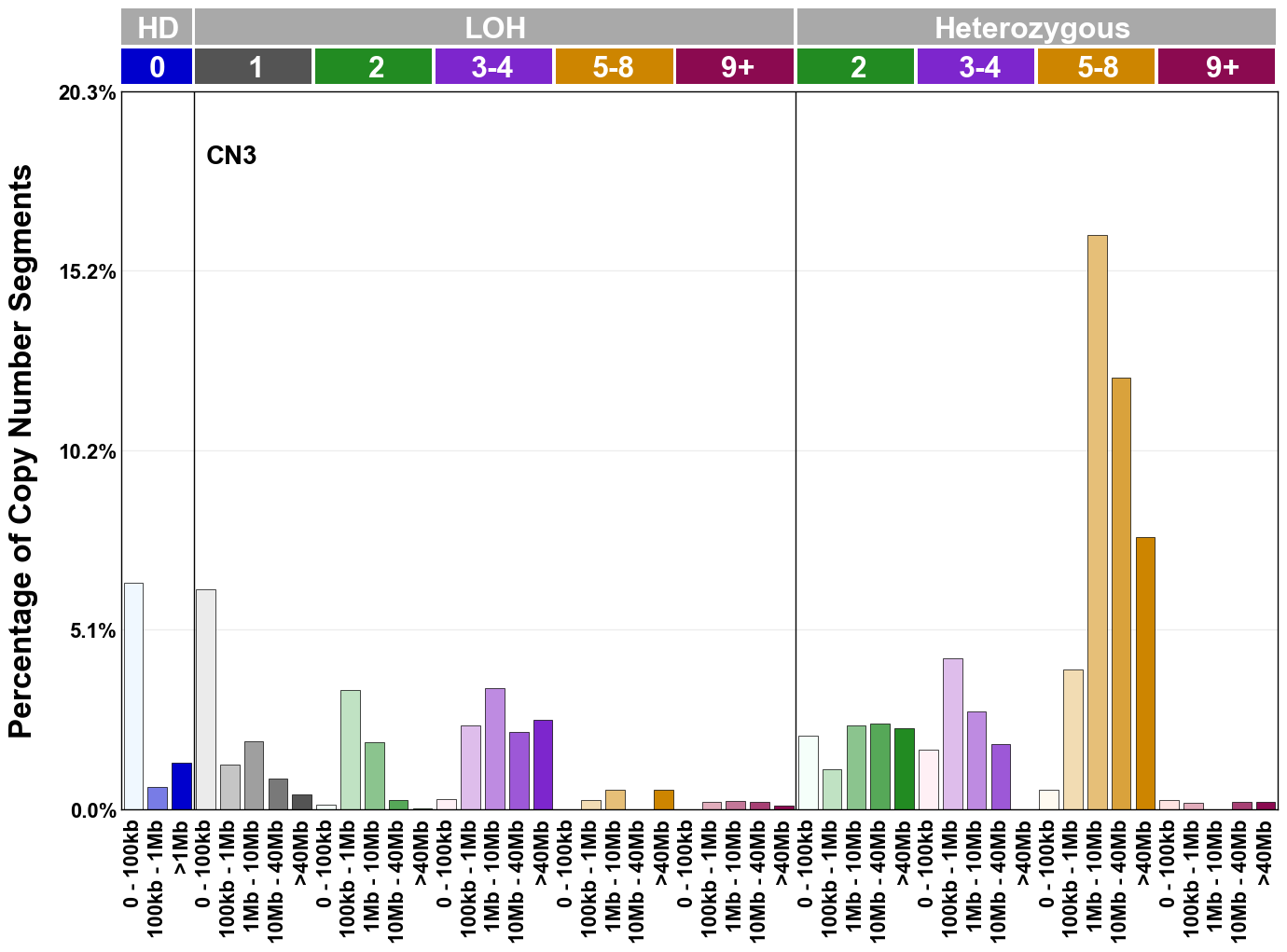
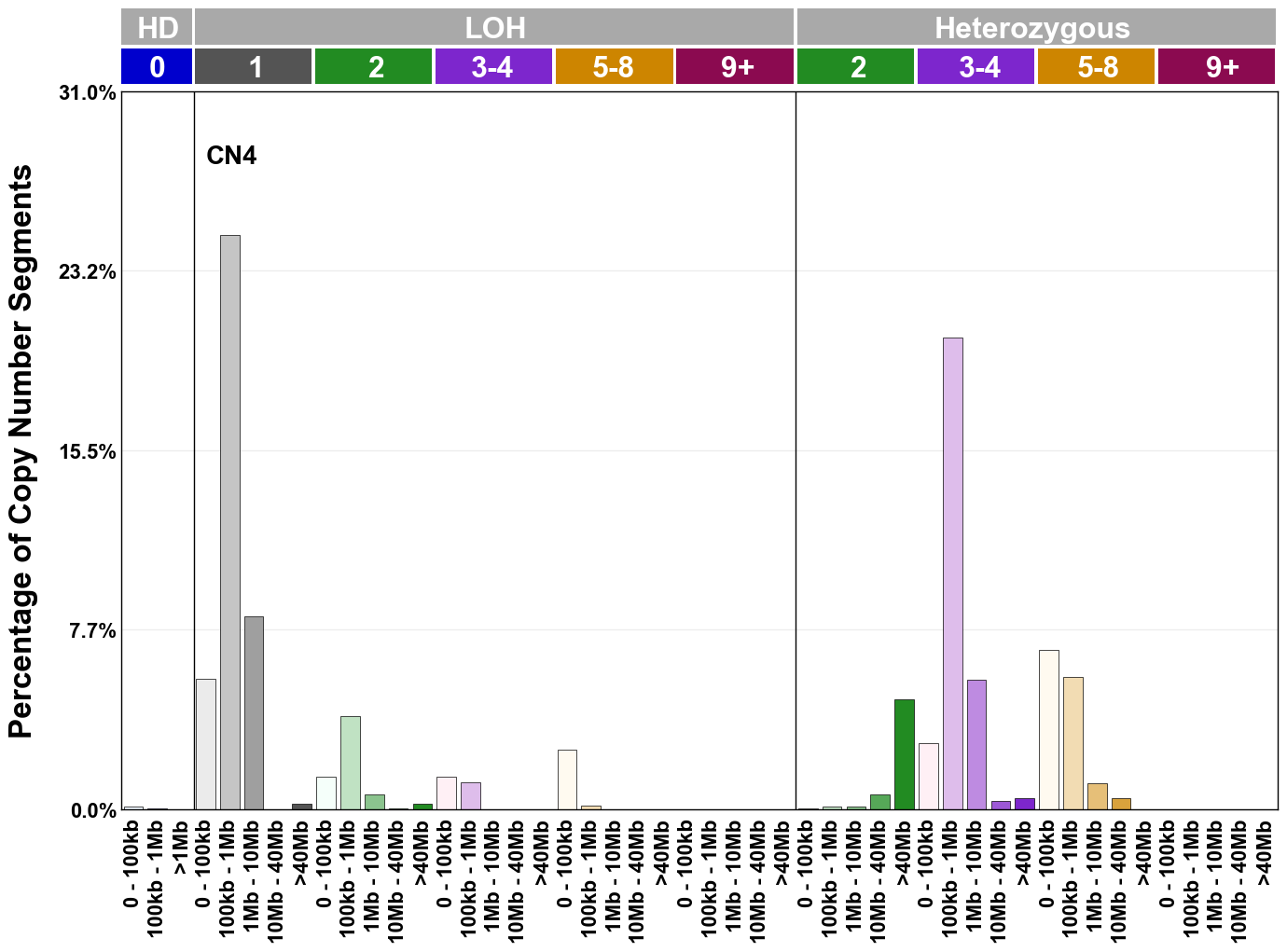
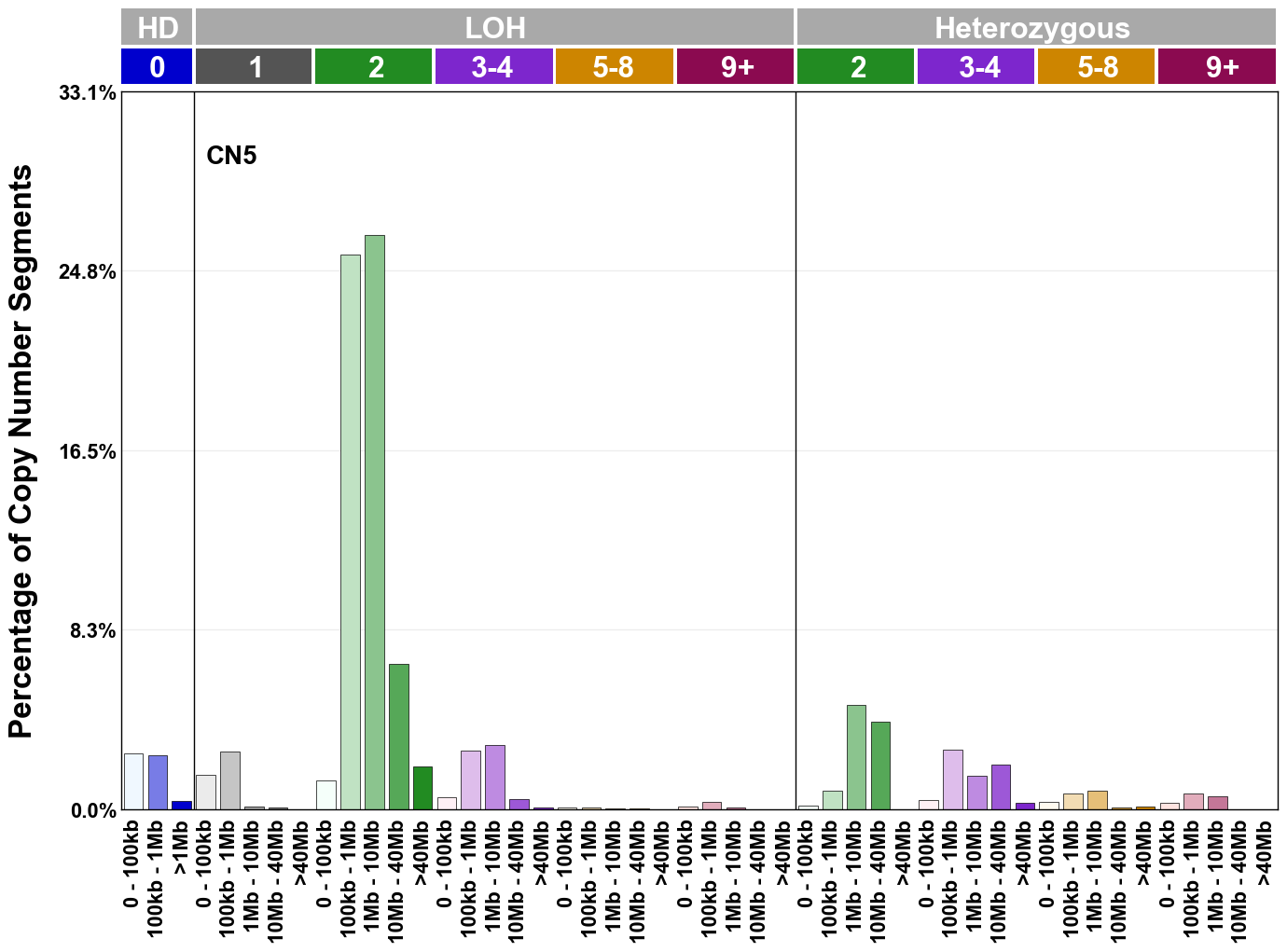
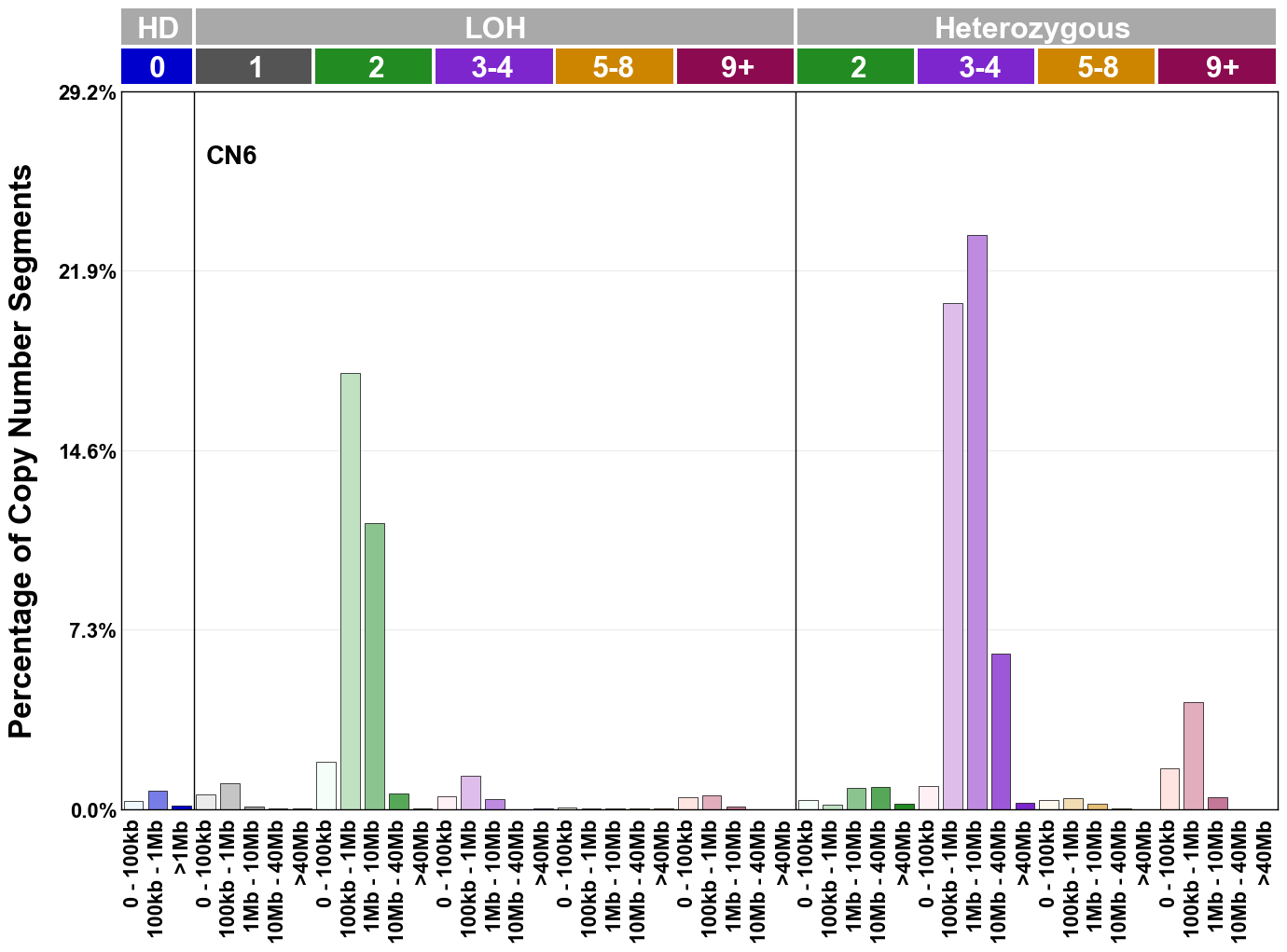
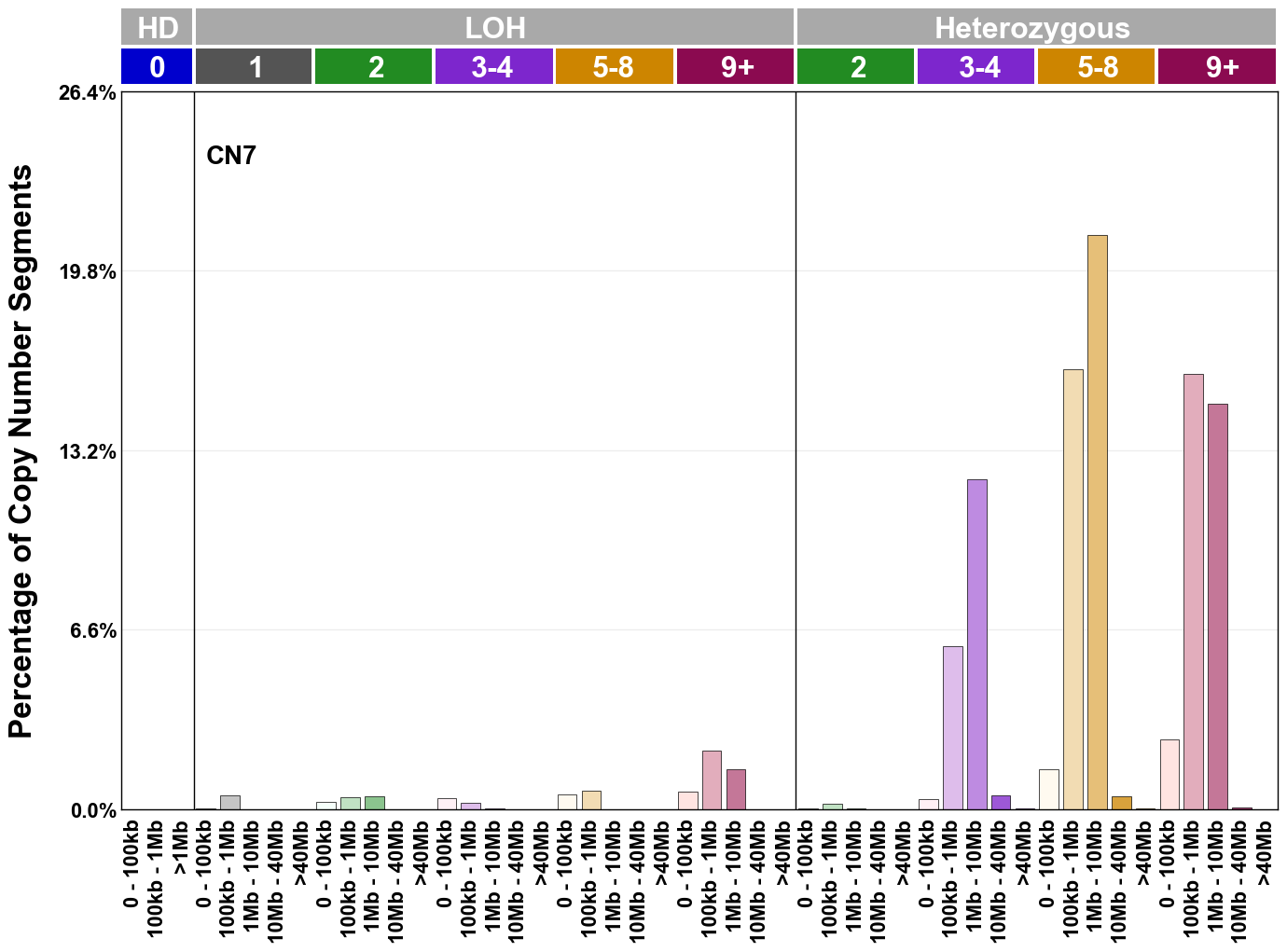
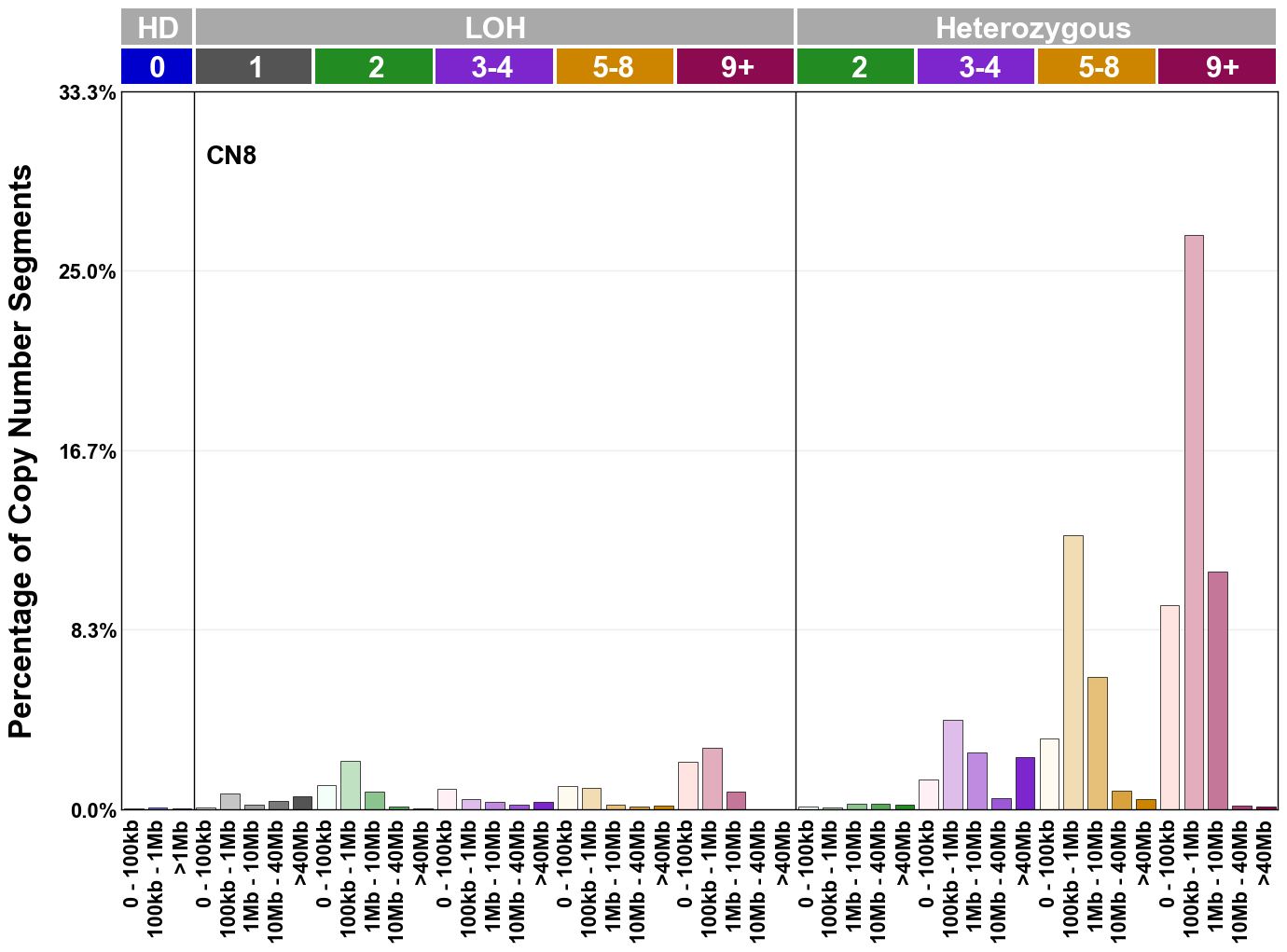
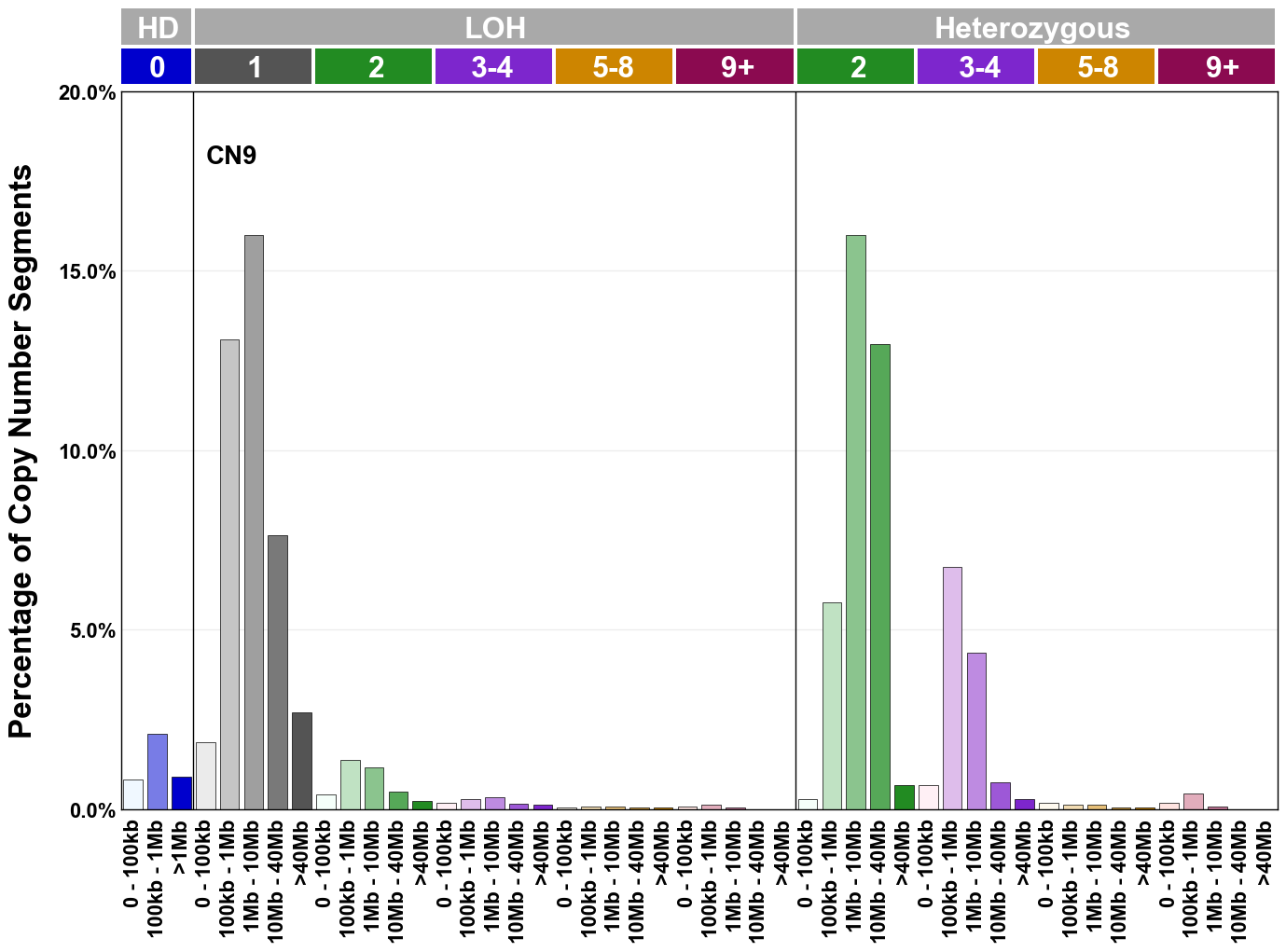
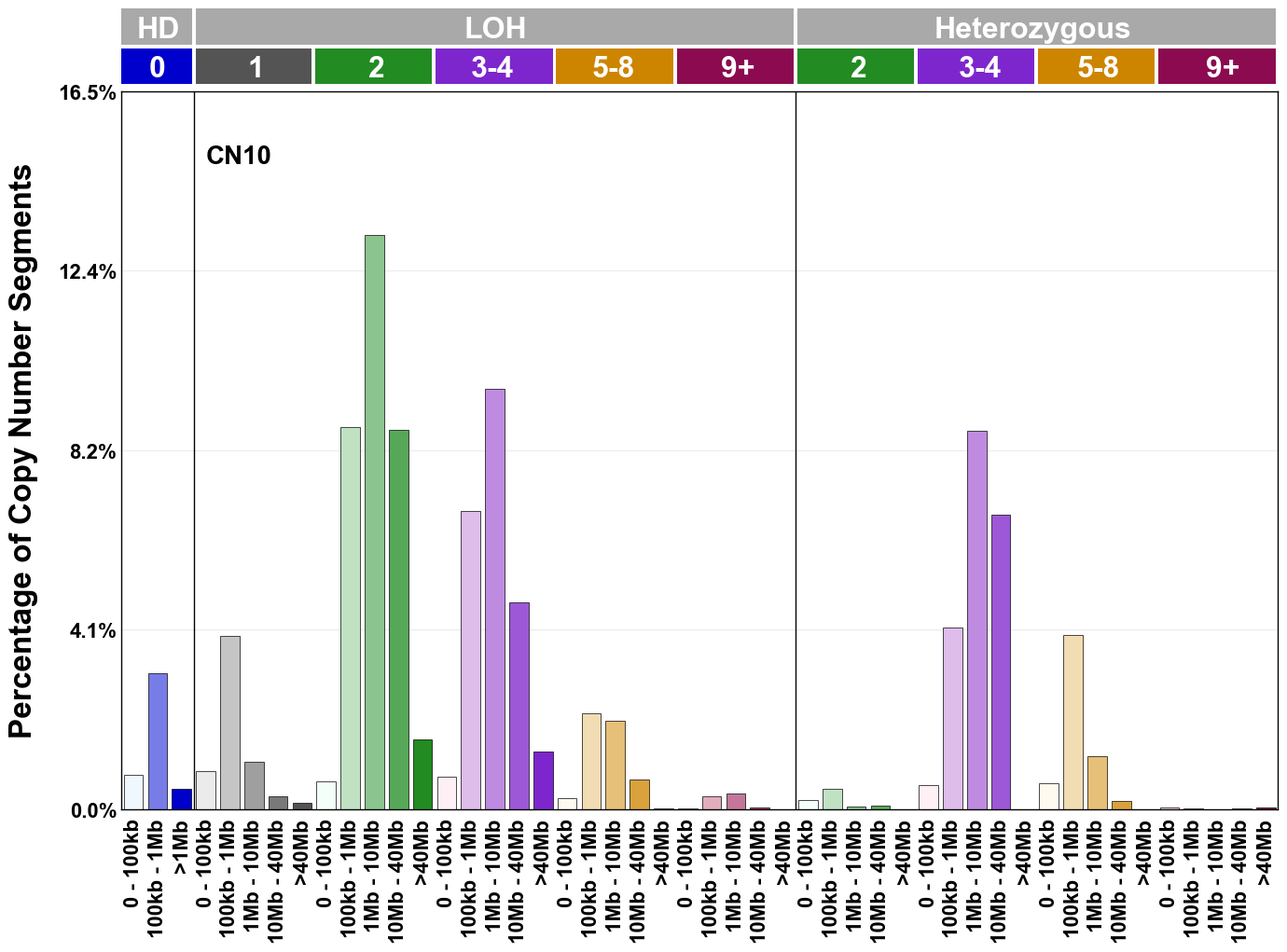
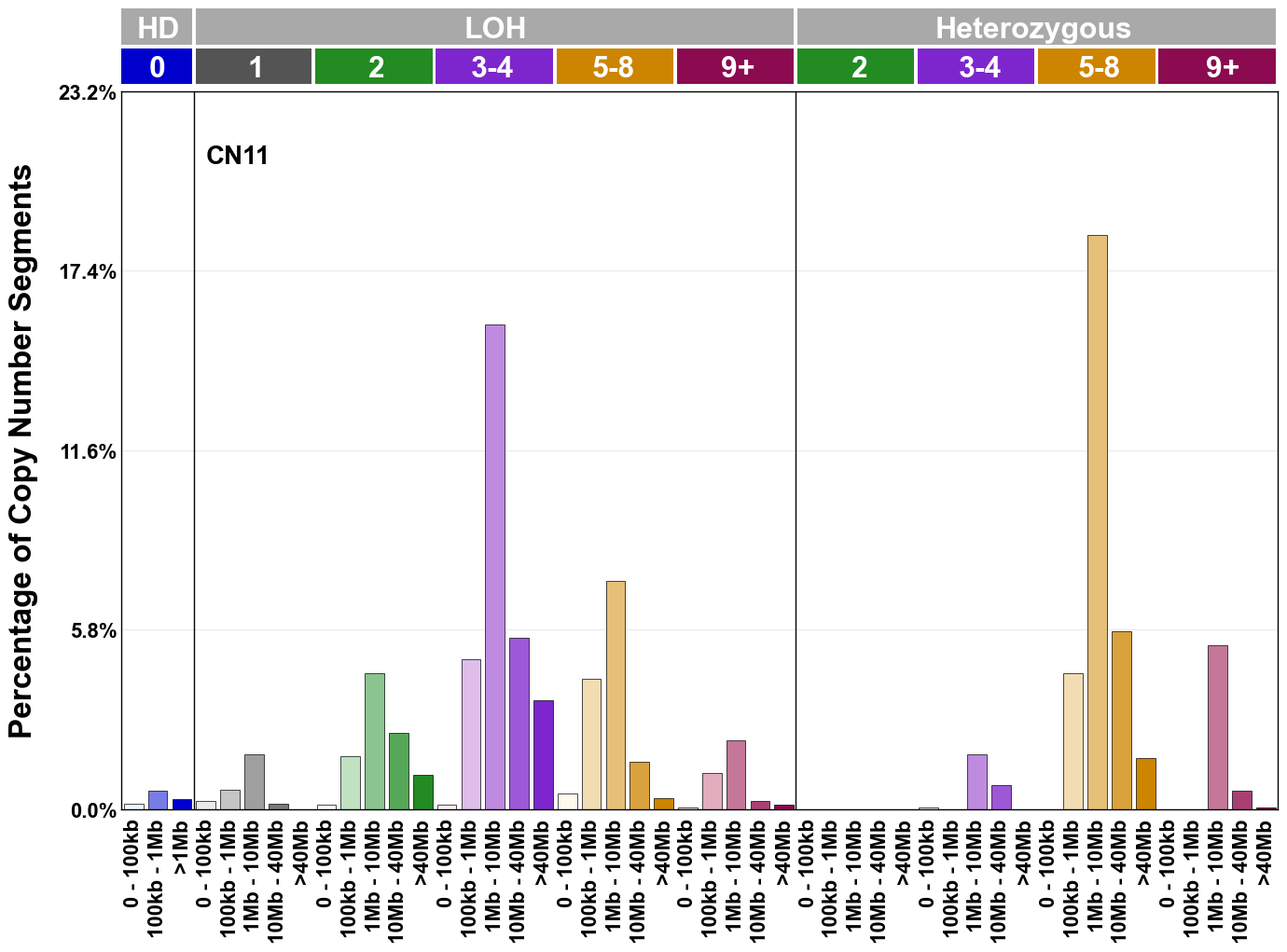
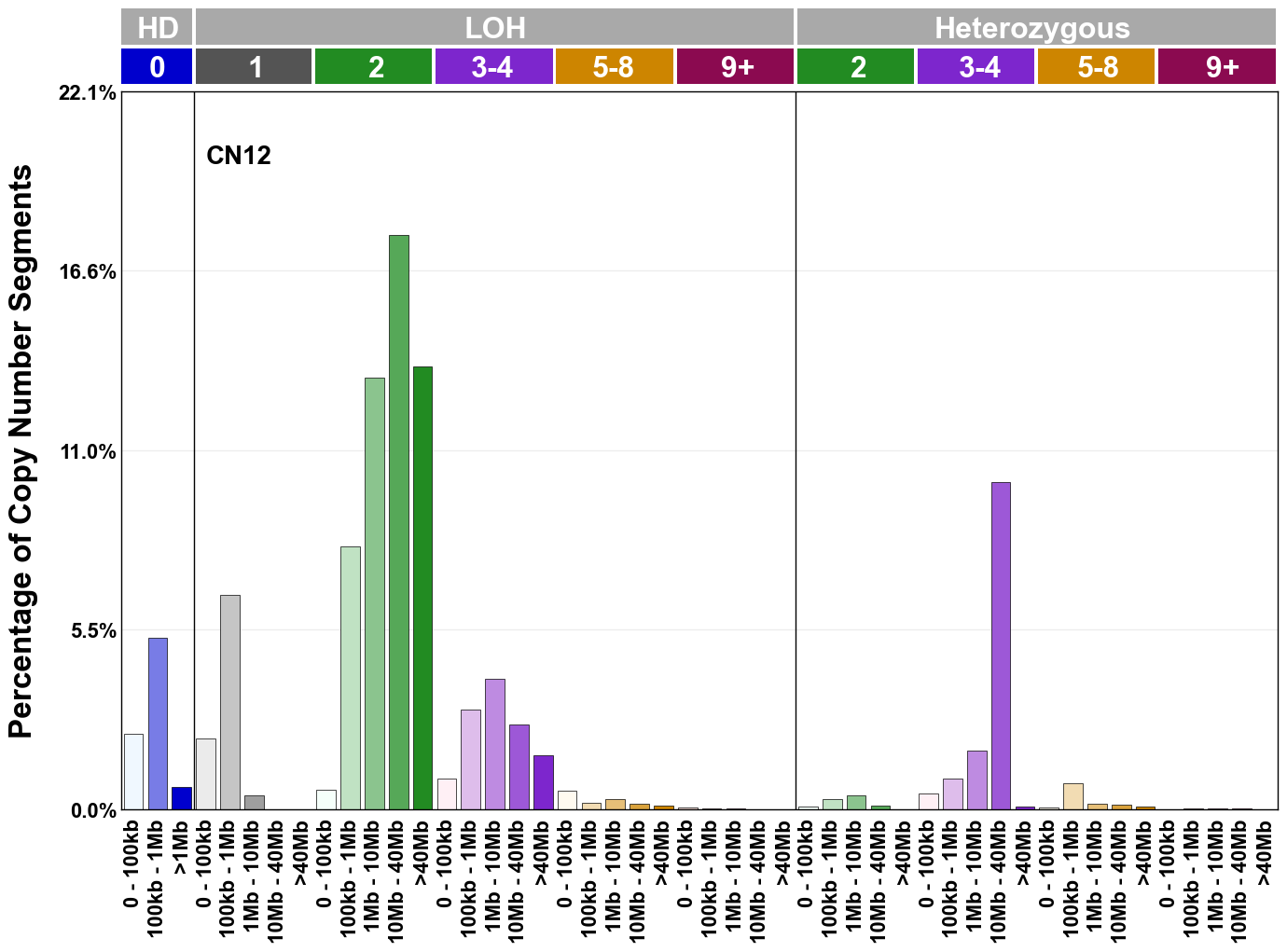
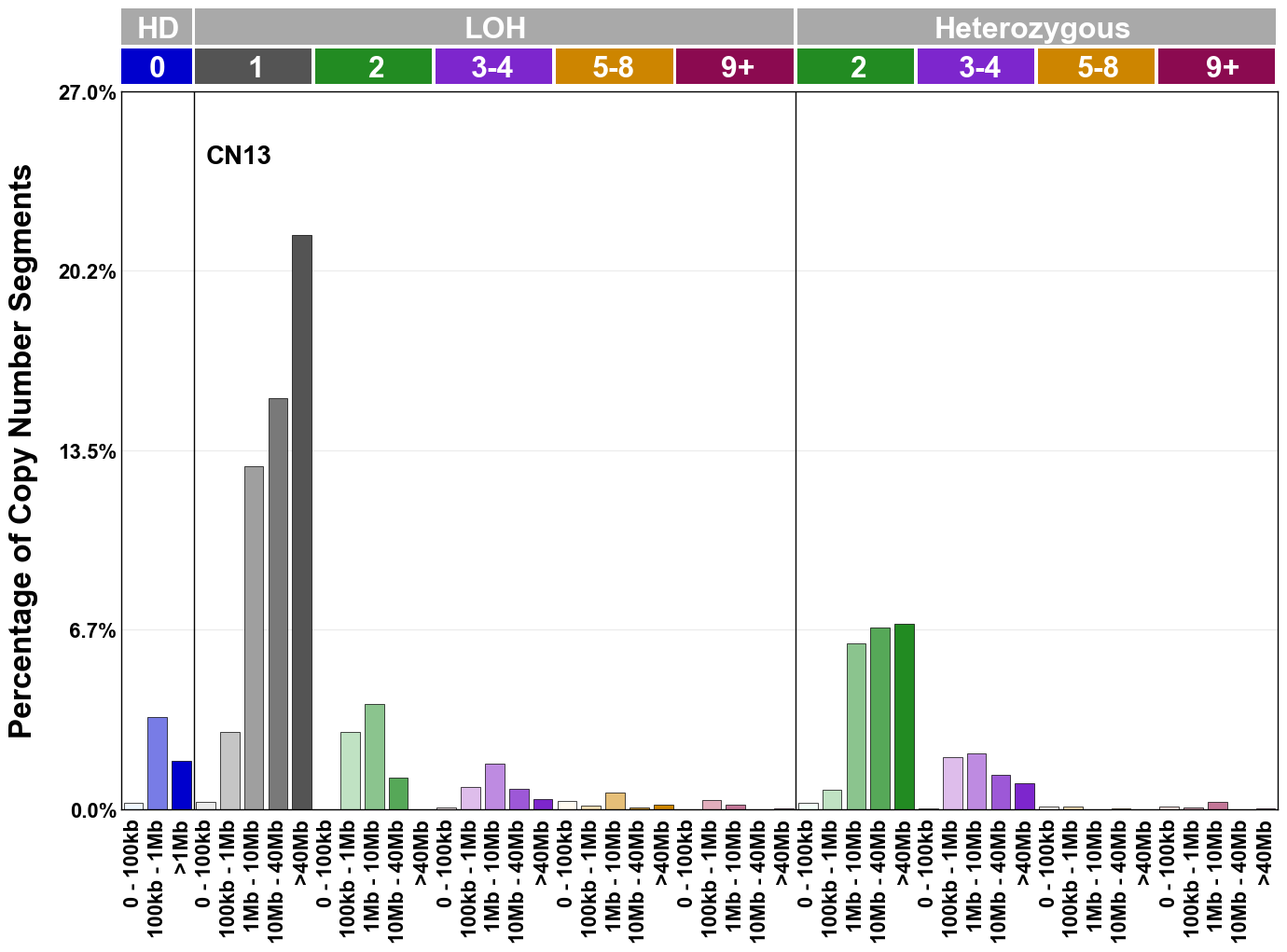
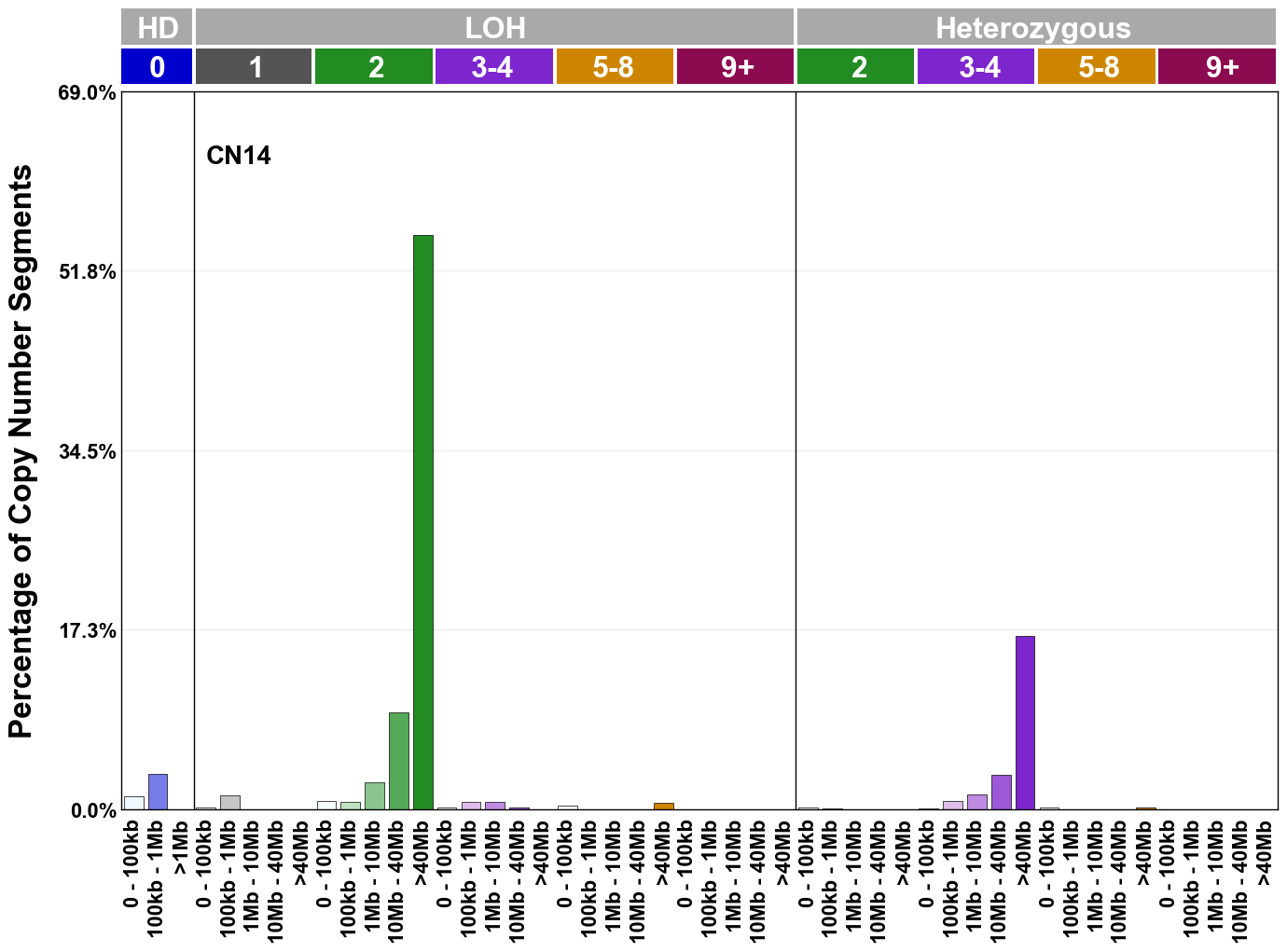
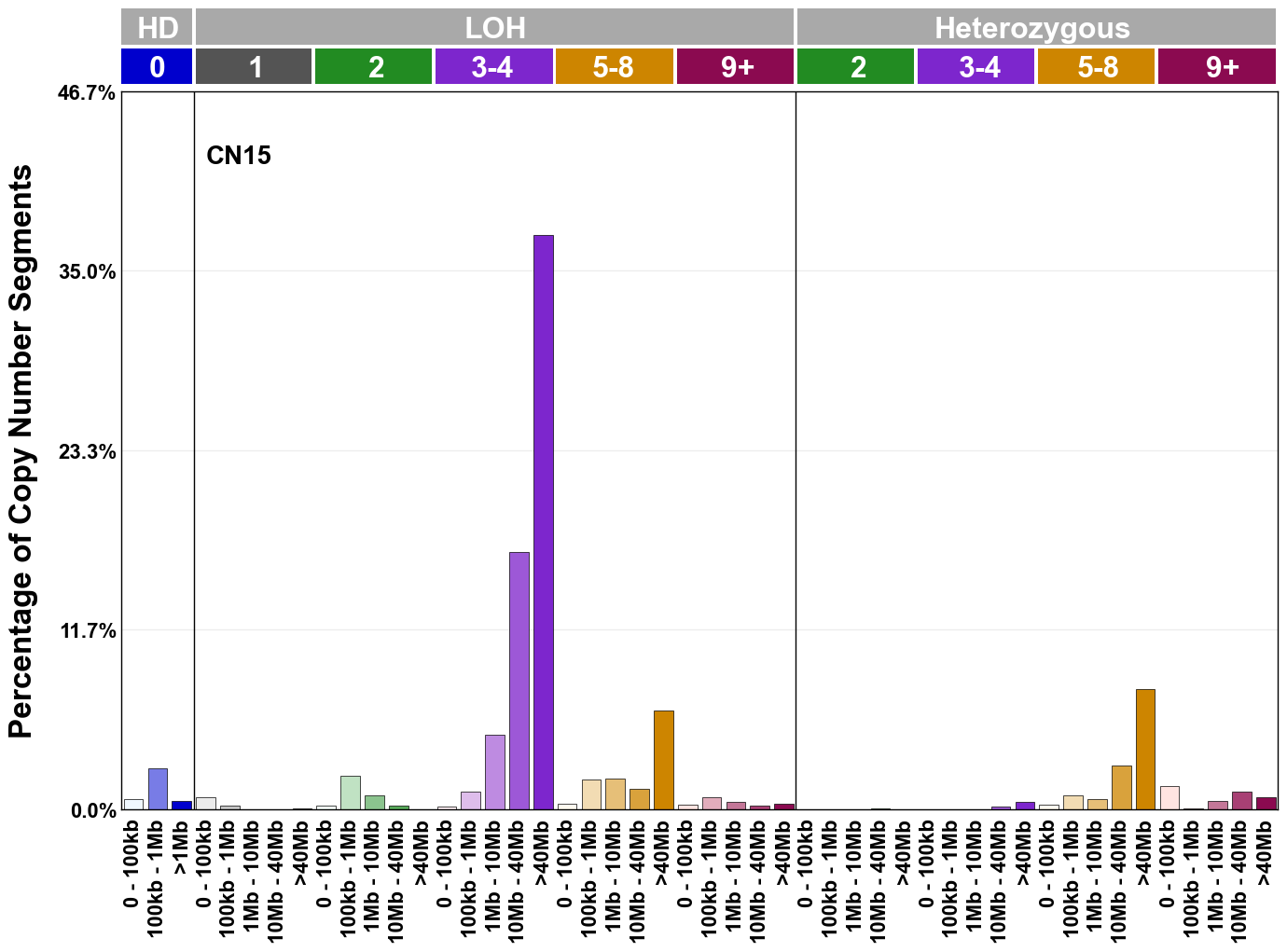
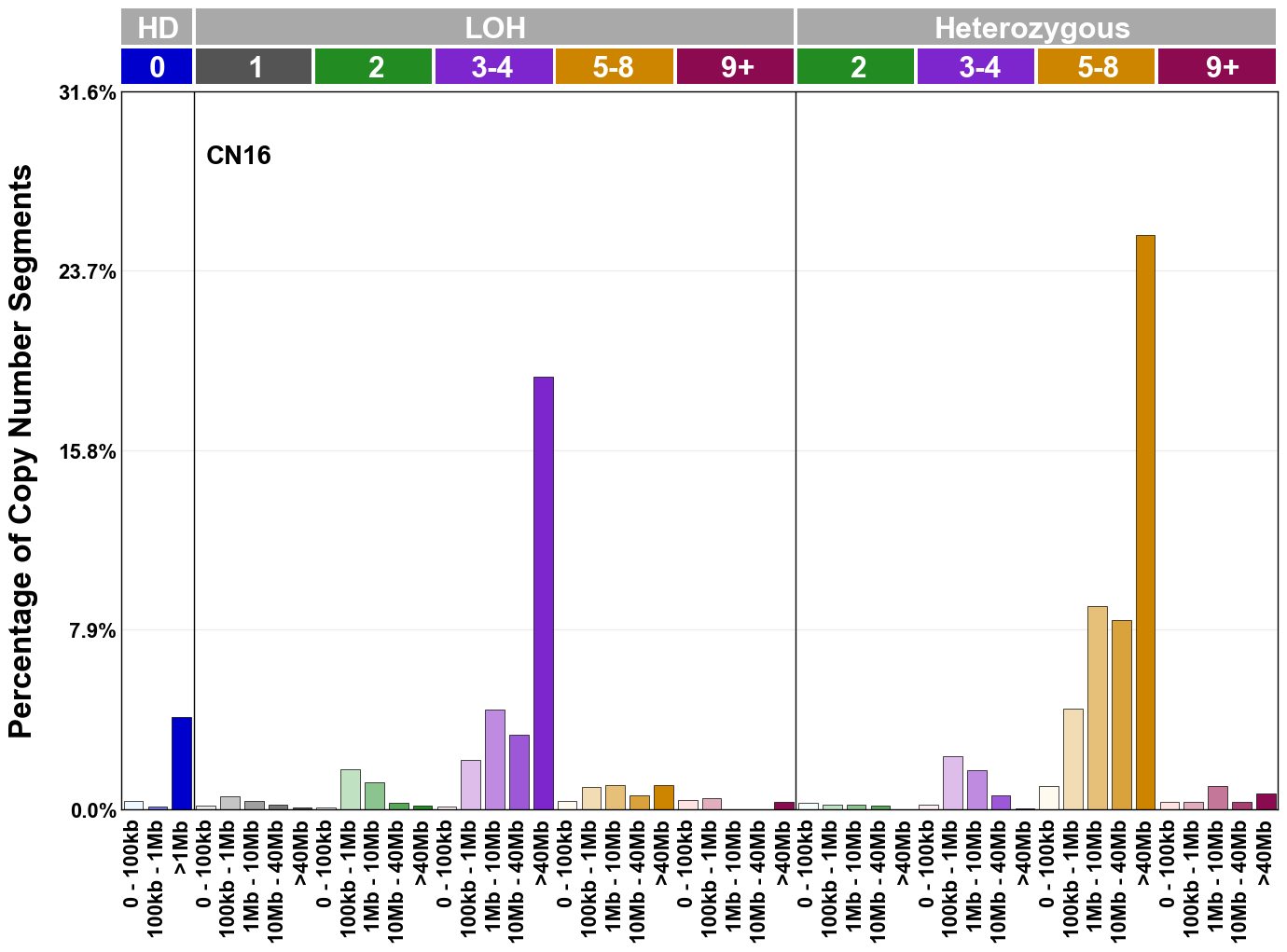
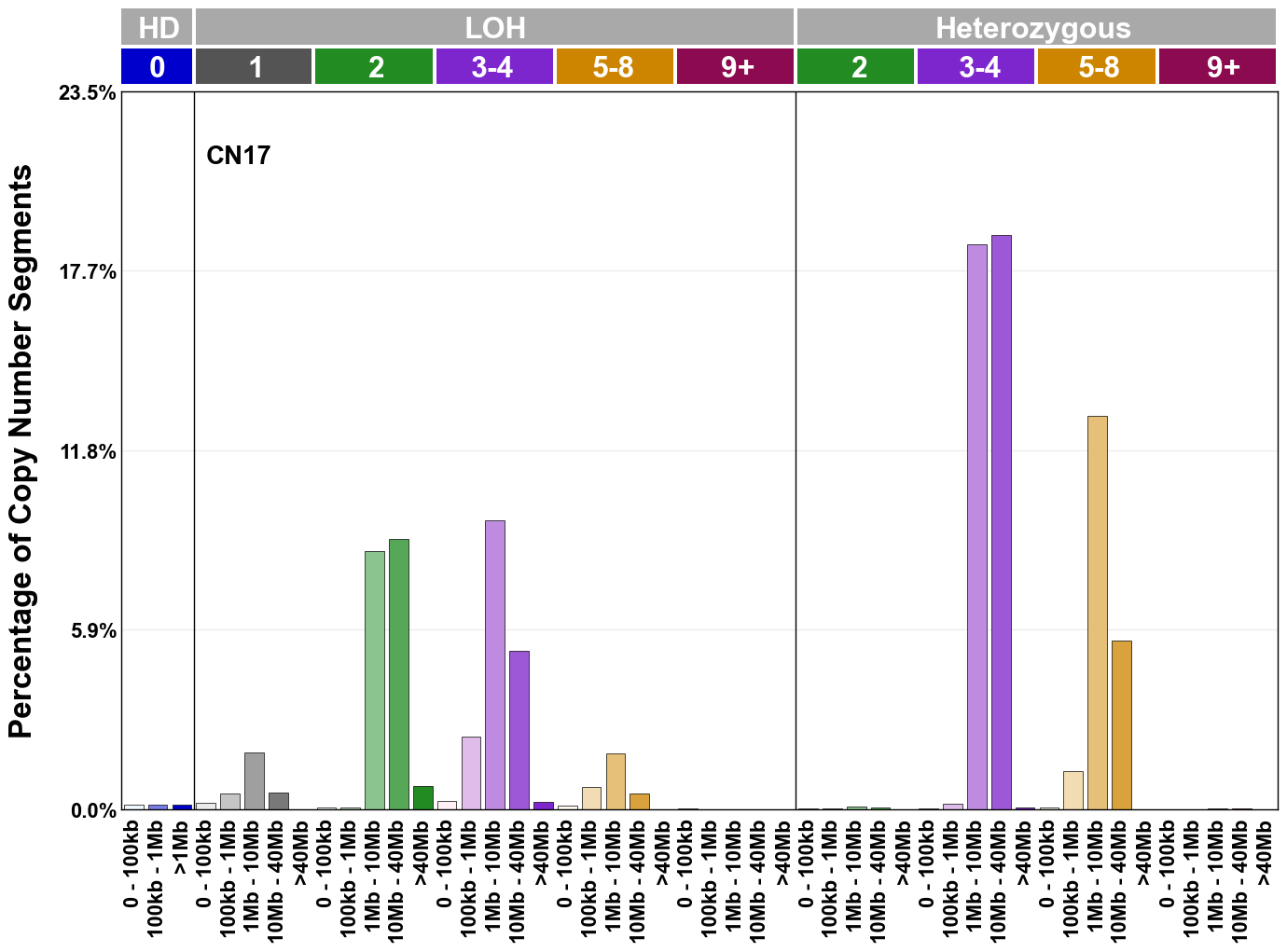
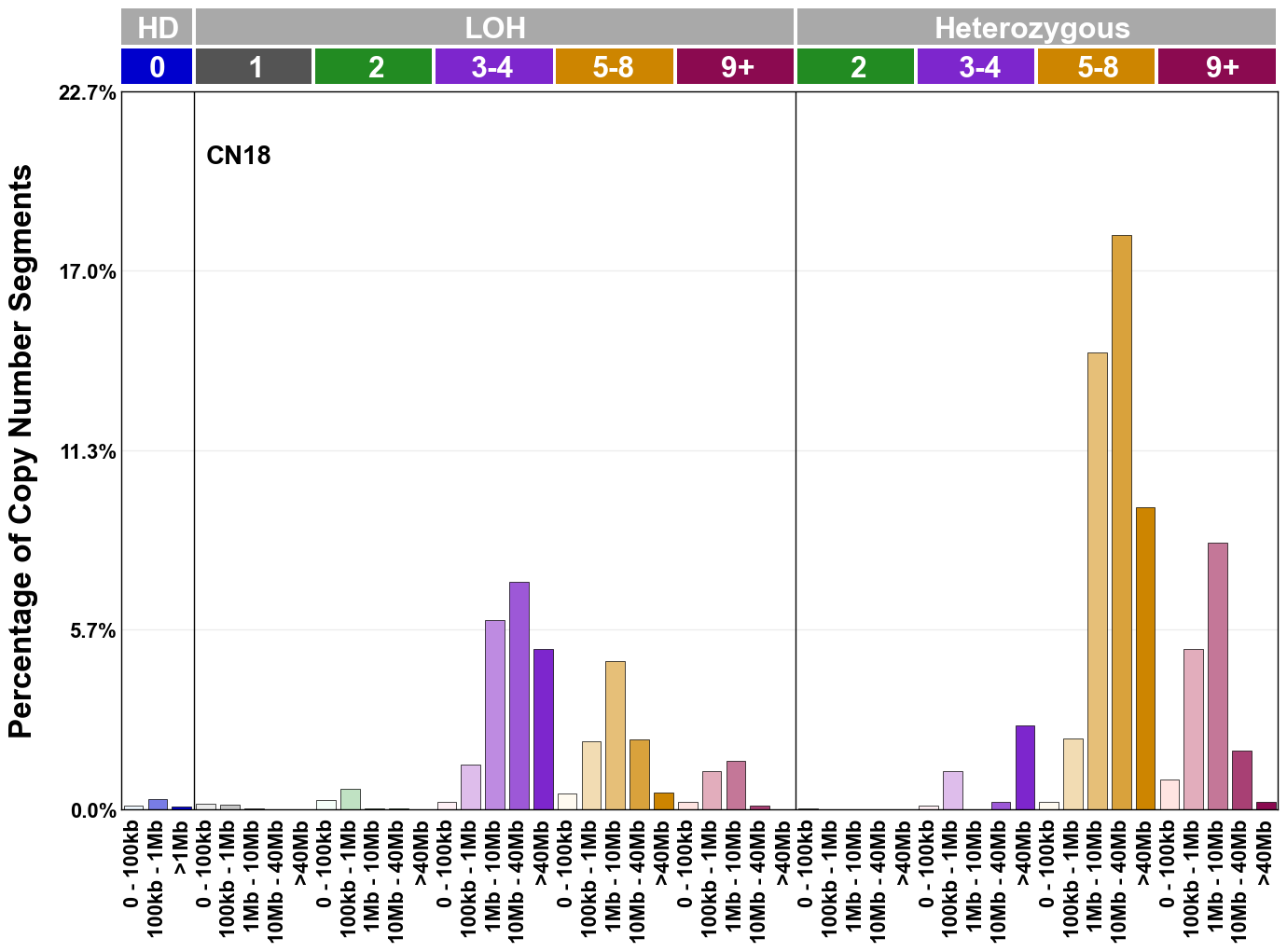
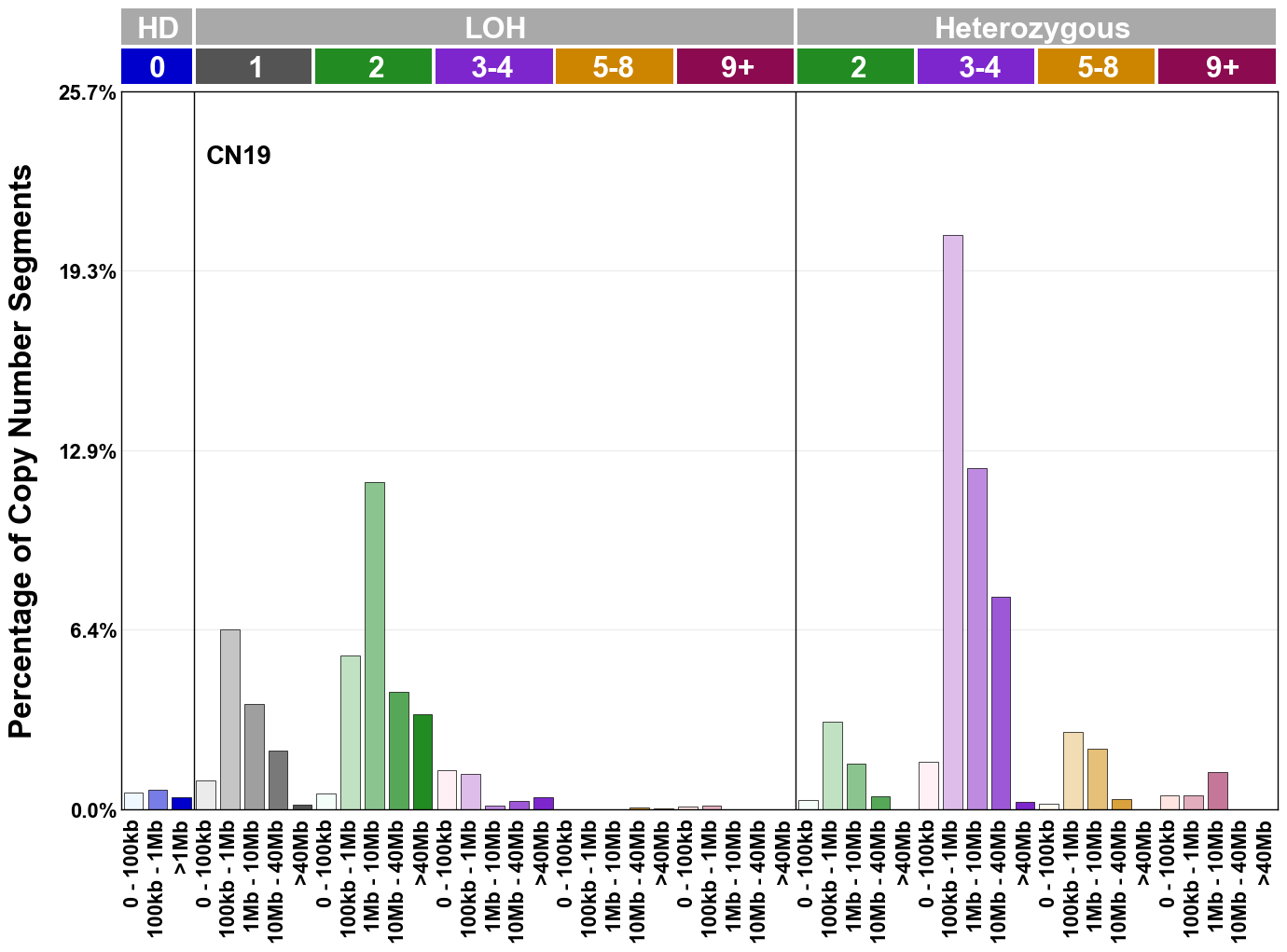
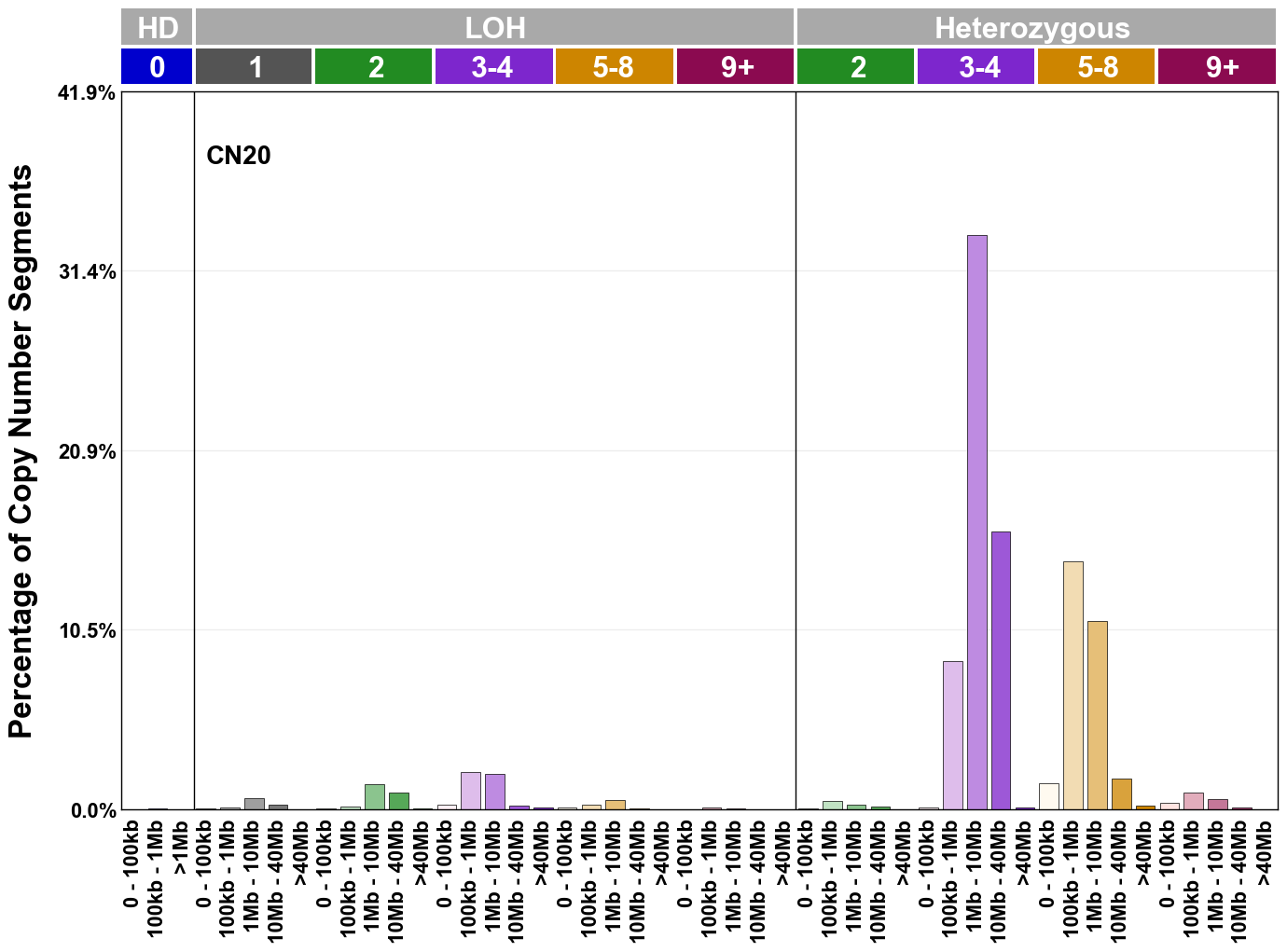
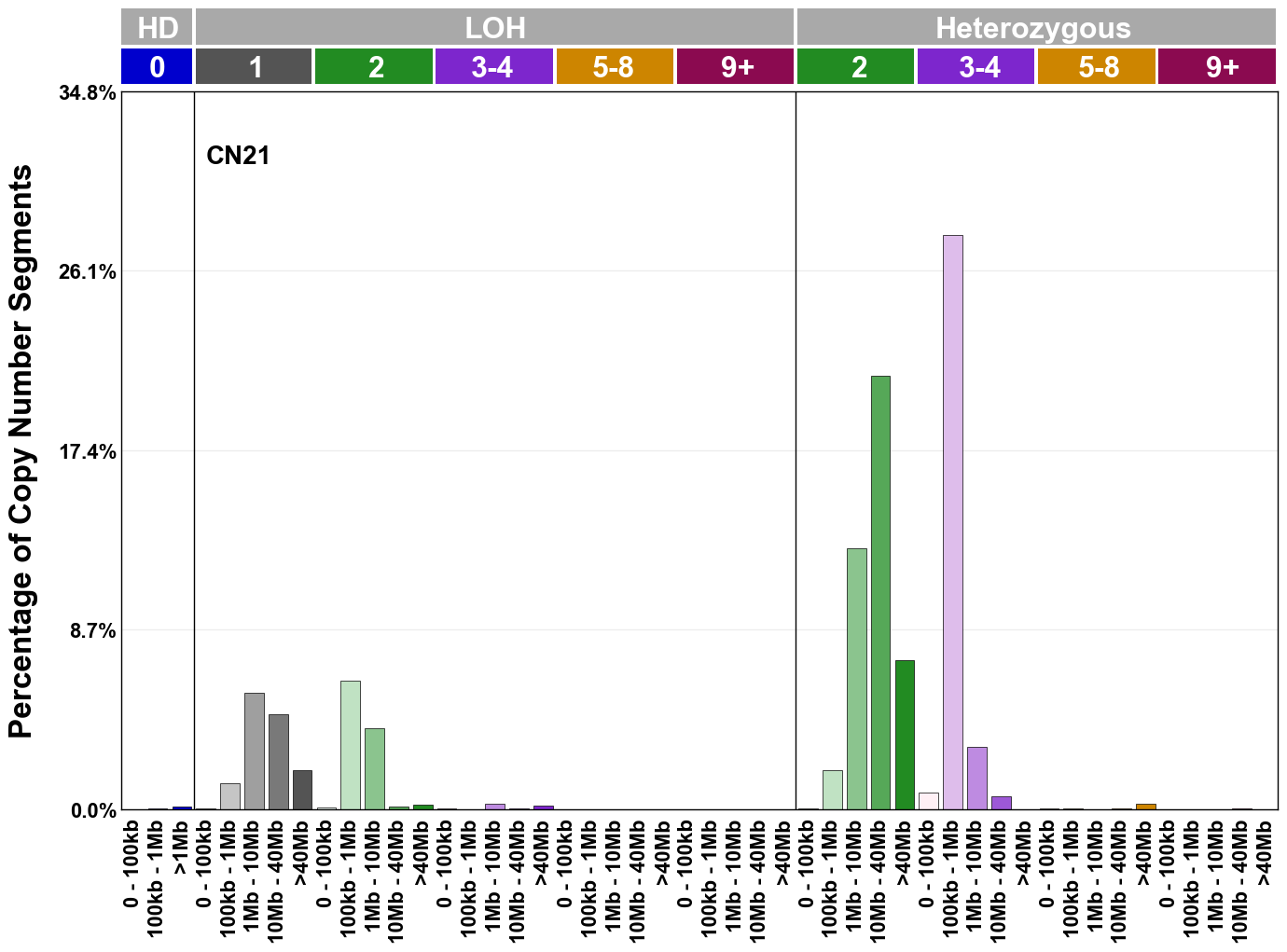
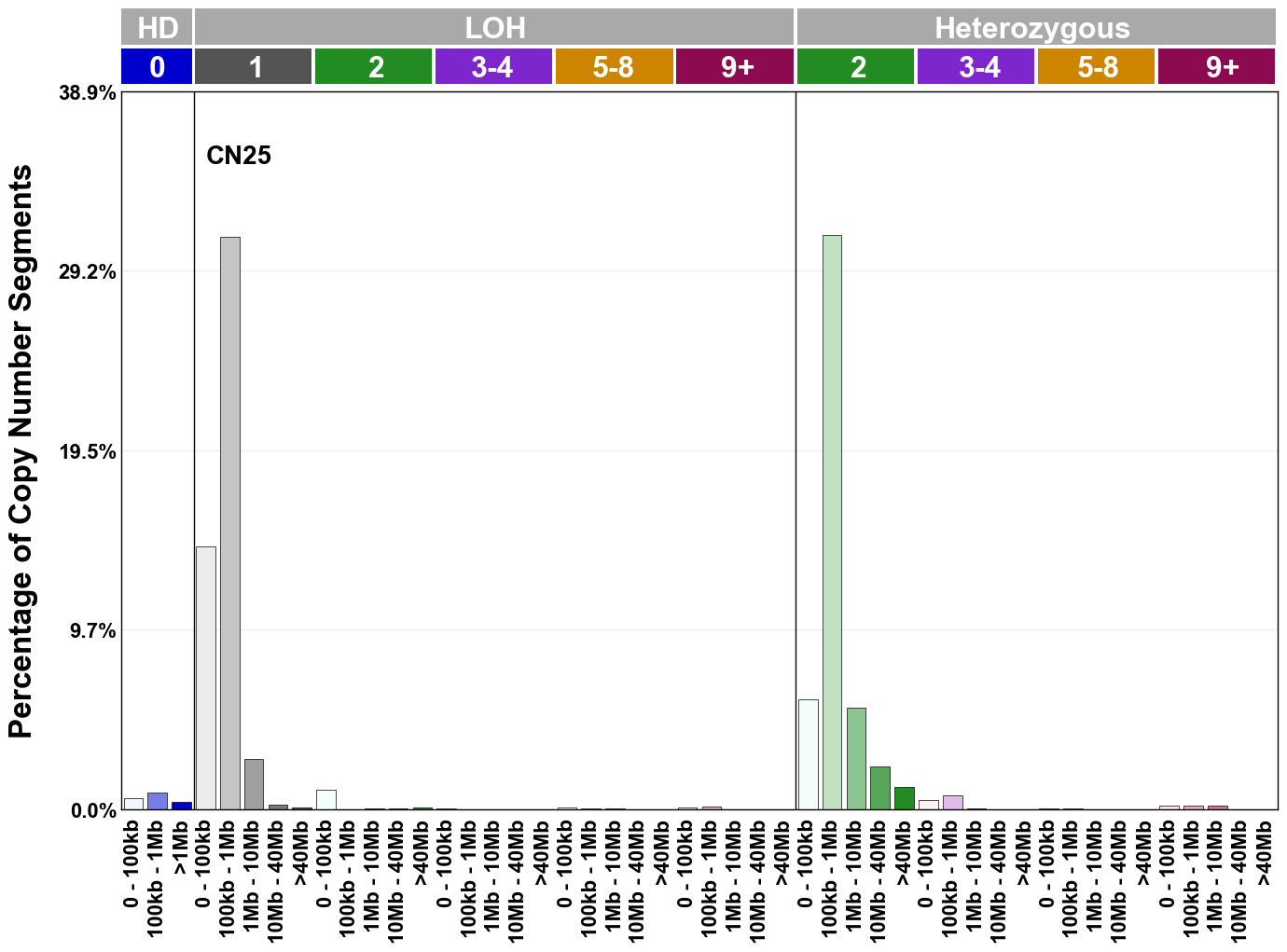
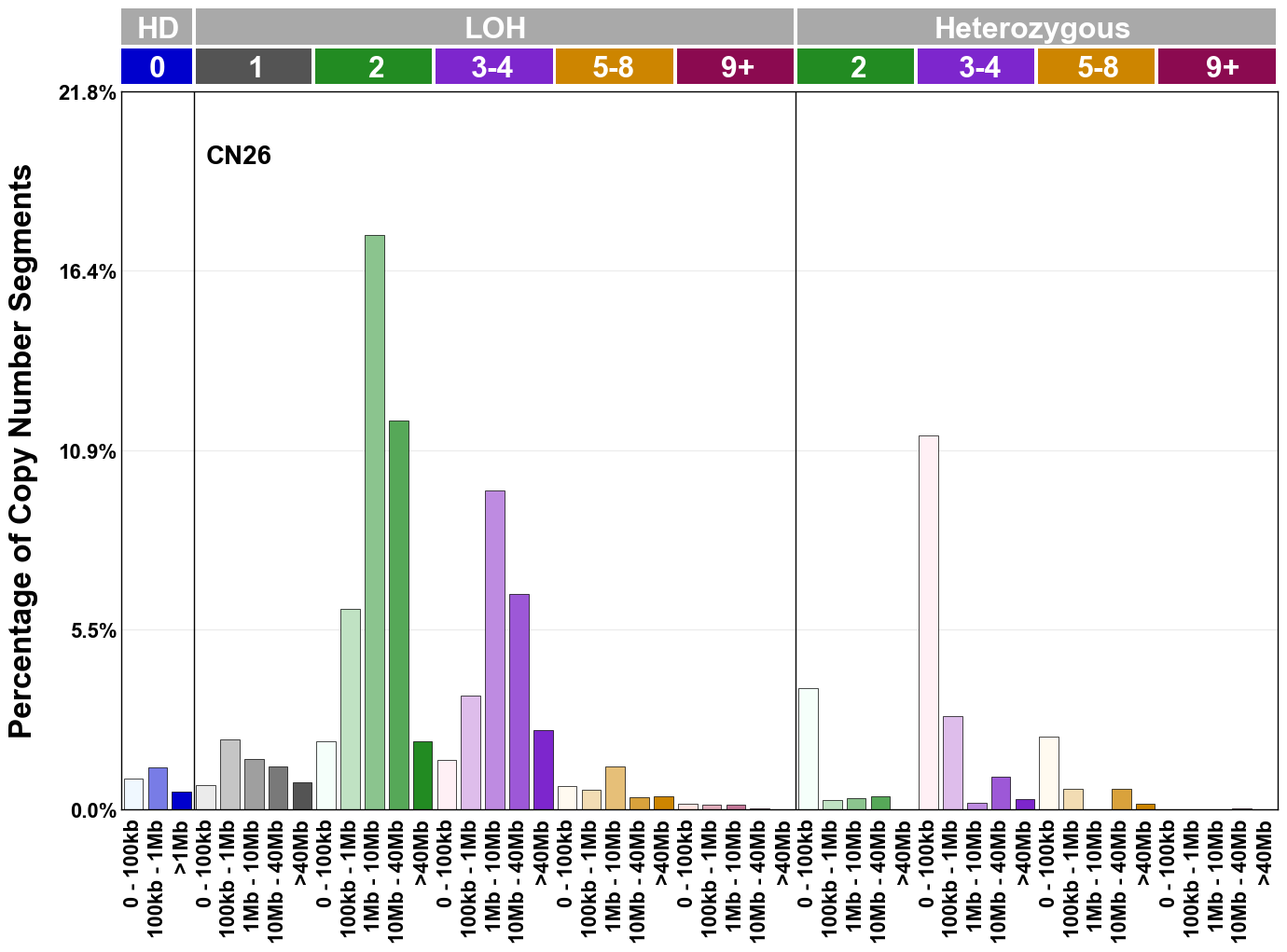
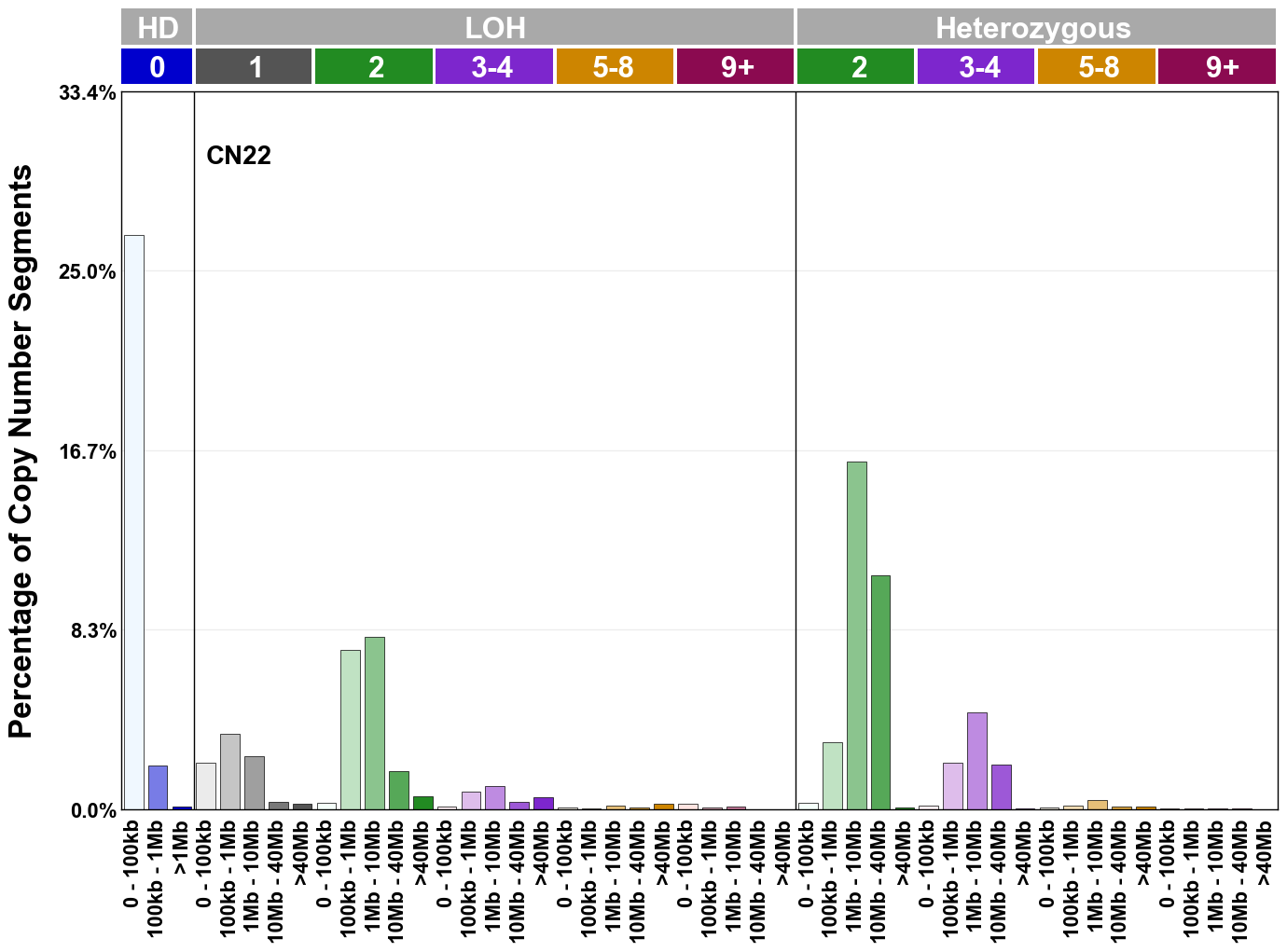
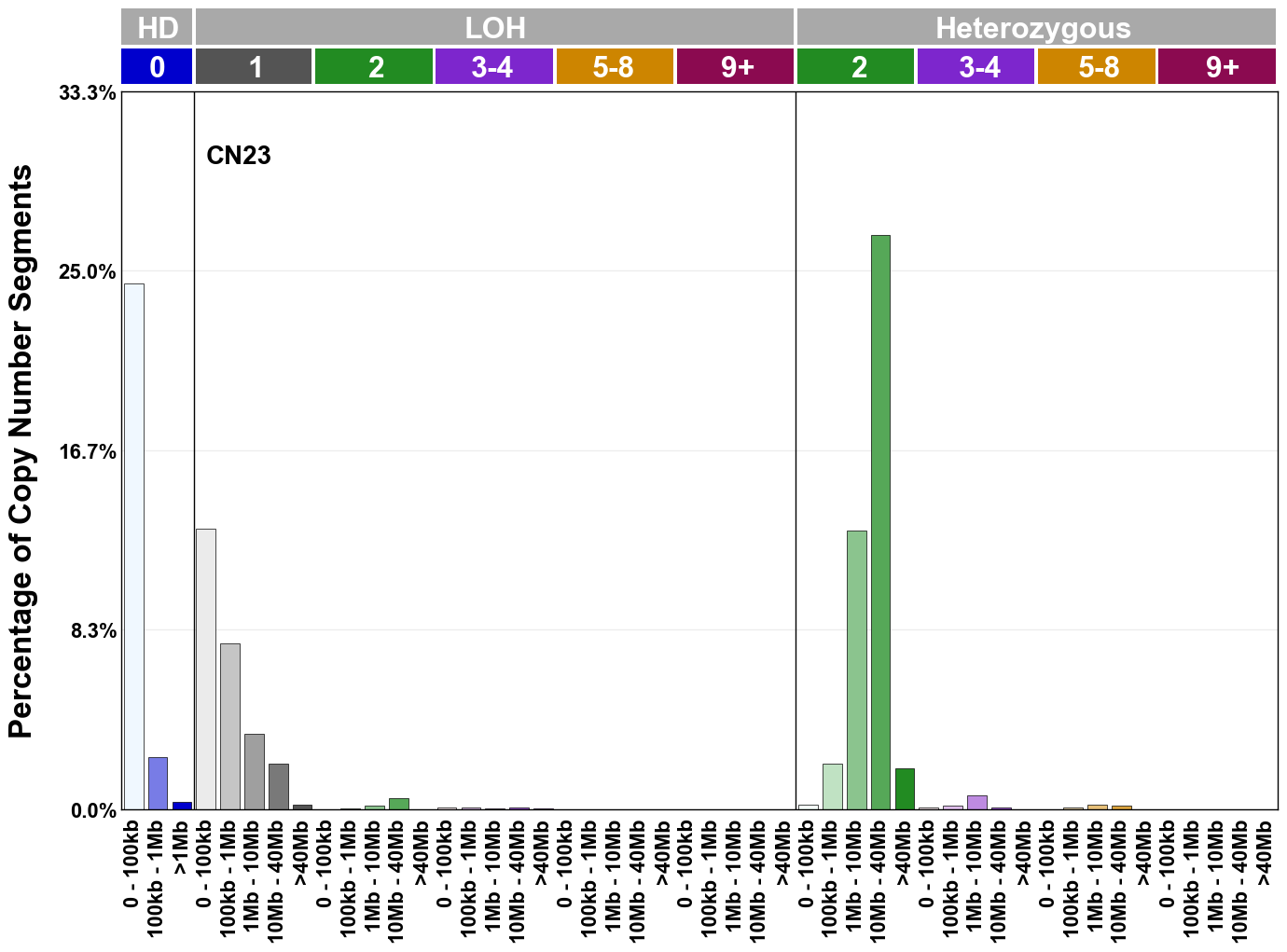
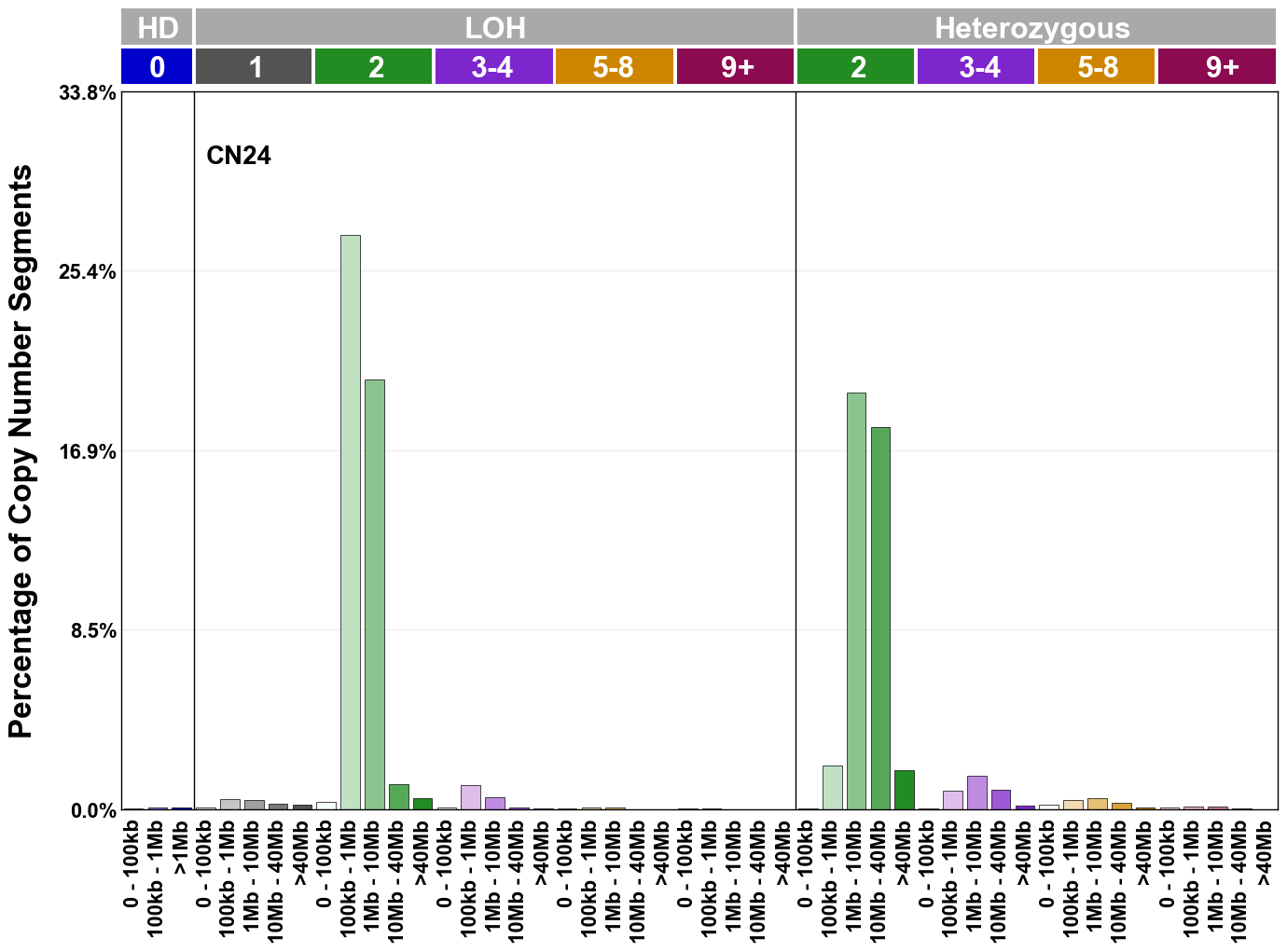
Copy number signatures are defined using the 48-channel copy number classification scheme. The scheme incorporates loss-of-heterozygosity status, total copy number state, and segment length to categorise segments from allele-specific copy number profiles (as major copy number and minor copy number respectively i.e. non-phased profiles), and the signatures displayed here were identified from 9,873 tumour copy number profiles obtained from The Cancer Genome Atlas (TCGA) SNP6 array data spanning 33 cancer types.
Click on any signature below to learn more about its details.
Methods
Signature extraction methods
The current set of reference signatures were extracted using sigProfiler (as described in Steele et al., 2022), from 9,873 allele-specific copy number profiles derived from SNP6 array data and called using ASCAT. These data were generated as part of The Cancer Genome Atlas project. Copy number profiles used to extract these signatures can be found here.
Further scripts for downstream analysis of copy number signatures can be found here and a discussion of copy number signatures can be found in this review.
COSMIC mutational signatures are available in numerical form in our data downloads page.