Mutational Signatures (v3.5 - November 2025)
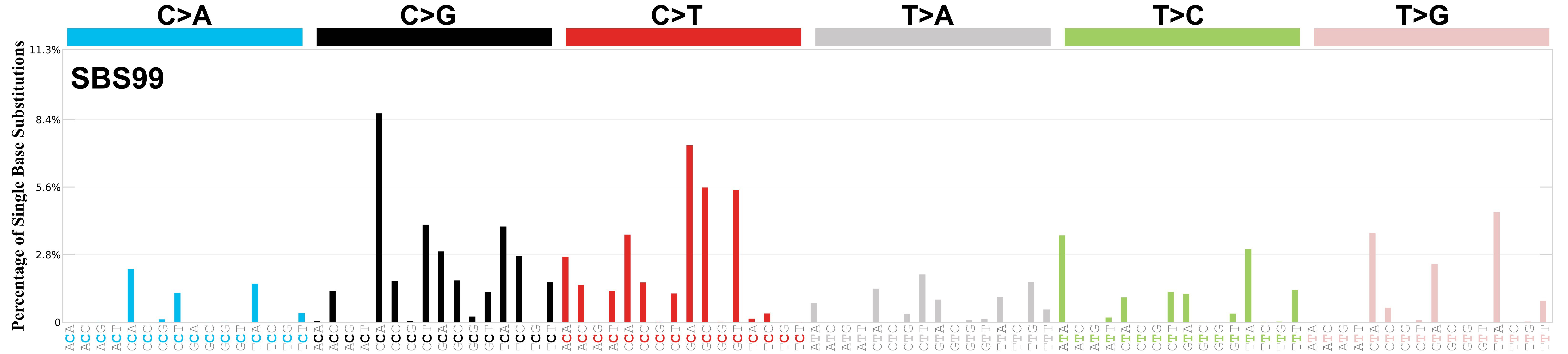
SBS99 · GRCh37 · COSMIC v103
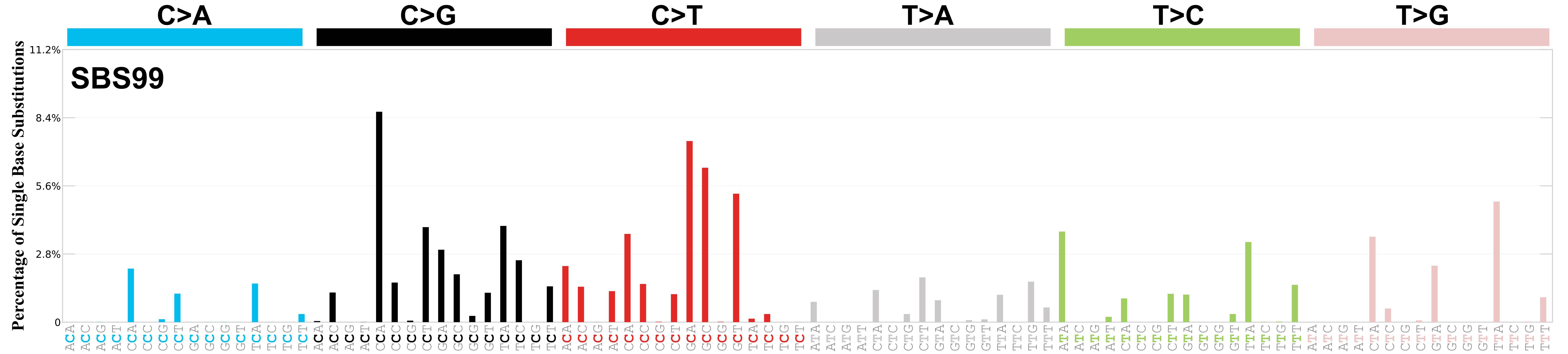
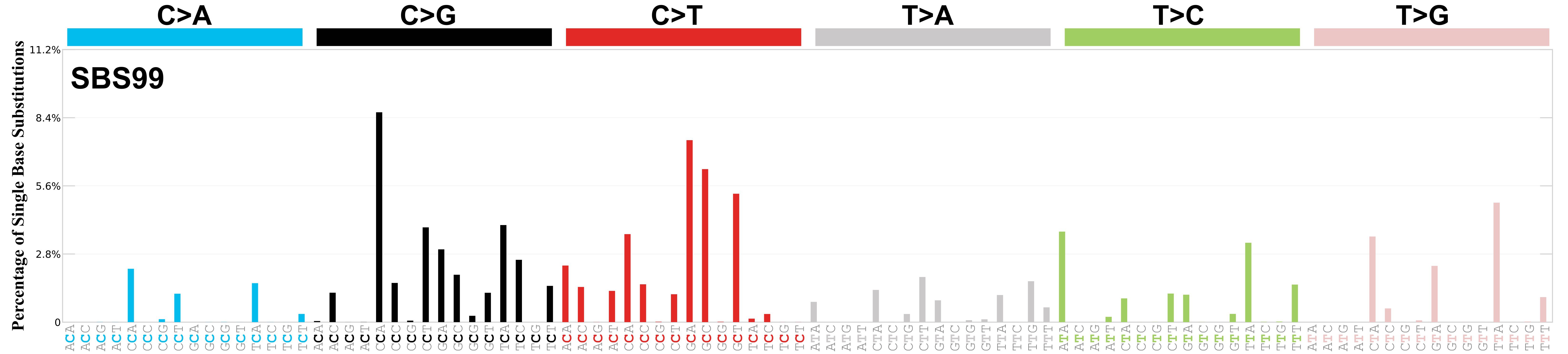
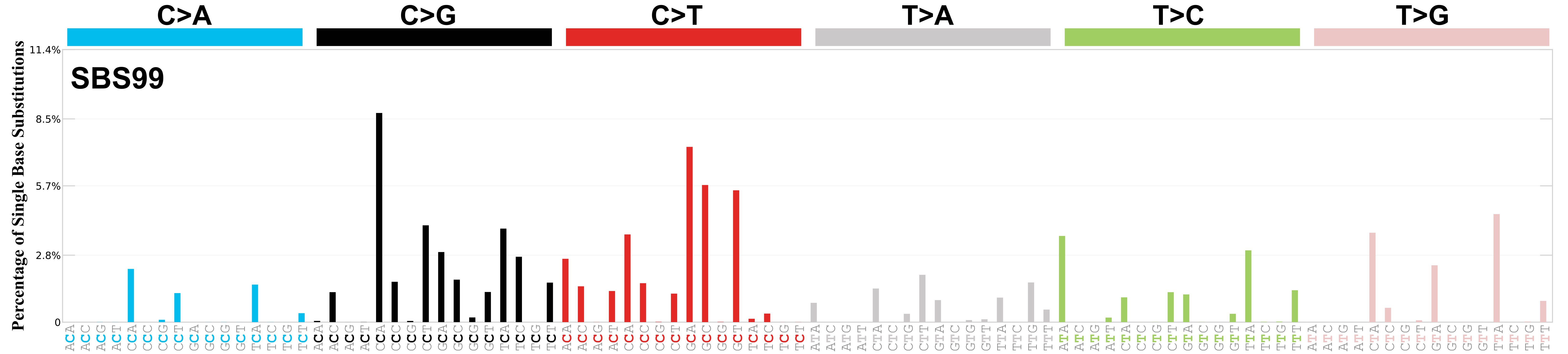
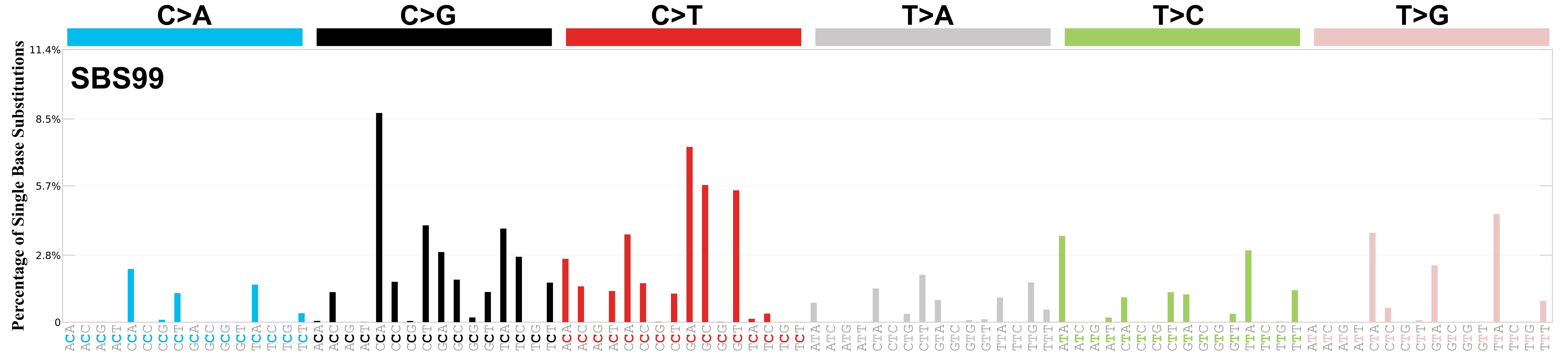
Mutational profile
Mutational profile using the conventional 96 mutation type classification. This classification is based on the six substitution subtypes: C>A, C>G, C>T, T>A, T>C, and T>G, as well as the nucleotides immediately 5’ and 3’ to the mutation.
Each of the substitutions is referred to by the pyrimidine of the mutated Watson—Crick base pair. Incorporating information on the bases immediately 5’ and 3’ to each mutated base generates 96 possible mutation types (6 types of substitution x 4 types of 5’ base x 4 types of 3’ base). Mutational signatures are displayed and reported based on the observed trinucleotide frequency of the genome, i.e., representing the relative proportions of mutations generated by each signature based on the actual trinucleotide frequencies of the corresponding reference genome.
Proposed aetiology
Exposure to melphalan treatment. Associations were found with patient samples that had prior exposure to melphalan and experimental validation was conducted on human cell lines with melphalan exposure.
Comments
Otherwise known as SBS-MM1 from Rustad et al. 2020 This signature was also detected in post-melphalan relapsed lymphomas (link to publication), and is associated with other post-melphalan therapy related myeloid and lymphoid signatures (link to publication). This signature was also found in a therapy-related bladder cancer in a patient previously exposed to melphalan for multiple myeloma (link to publication)
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Rustad et al. 2020 Nature Communications | v3.4 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WES & WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | Yes | Strong transcriptional strand bias in C>T | |
| Proposed aetiology | Mutational process | Support | |
| Melphalan exposure | Experimental confirmation | ||
| Experimental validation | Experimental study | Species | |
| Rustad et al. 2020 Nature Communications | Human cell lines | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Found in multiple myeloma and certain lymphomas.