Mutational Signatures (v3.6 - May 2026)
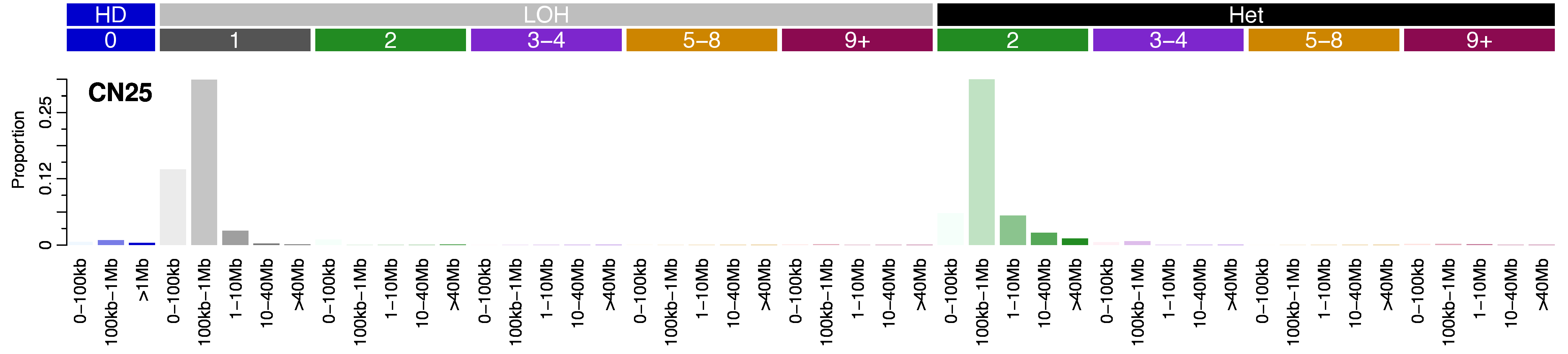
CN25 · GRCh37 · COSMIC v104
Mutational profile
Copy number signatures are defined by a 48 context copy number classification scheme. The scheme incorporates loss-of-heterozygosity status, total copy number state, and segment length to categorise segments from allele-specific copy number profiles.
The scheme corresponds to three heterozygosity states:
- heterozygous segments with copy number of [A>0, B>0] (numbers reflect the counts for major allele A and minor allele B)
- segments with LOH with copy number of [A>0, B=0]
- segments with homozygous deletions [A=0, B=0]
Segments are further subclassified into 6 classes based on the sum of major and minor allele (total copy number) and chosen for biological relevance:
- total copy number (TCN)=0 (homozygous deletion)
- TCN=1 (deletion leading to LOH)
- TCN=2 (wild type, including copy-neutral LOH)
- TCN=3 or 4 (minor gain)
- TCN=5 to 8 (moderate gain)
- TCN>=9 (high-level amplification).
Each of the heterozygous and LOH total copy number states are then organised into five classes based on the size of their segments: 0 – 100kb, 100kb – 1Mb, 1Mb – 10Mb, 10Mb – 40Mb, and >40Mb (the largest category for homozygous deletions was restricted to >1Mb) in order to capture focal, large scale, and chromosomal-scale copy number changes. This gives a total of 48 copy number context to categorise segments into (3x homozygous deletion categories, 5*5=25x LOH categories, 4*5=20 heterozygous categories).
Proposed aetiology
CN25 is associated with MSH6 inactivation and dMMR.
Comments
CN25 is composed of <1Mb loss of heterozygosity (LOH) deletions (similar to CN9 involving LoH >1Mb), which is indicative of chromothripsis. However, unlike other chromothripsis-associated CN signatures, CN4 and CN8, CN25 is composed of low total copy number states with single copy LOH and diploid heterozygous states.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Everall et al. 2026 Nature Genetics | v3.4 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | No | - | |
| Proposed aetiology | Mutational process | Support | |
| MMR deficiency | Statistical association | ||
| Experimental validation | Experimental study | Species | |
| - | - | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Feature of CNS, connective tissue, and prostate cancers.