Mutational Signatures (v3.6 - May 2026)
CN6 · GRCh37 · COSMIC v104
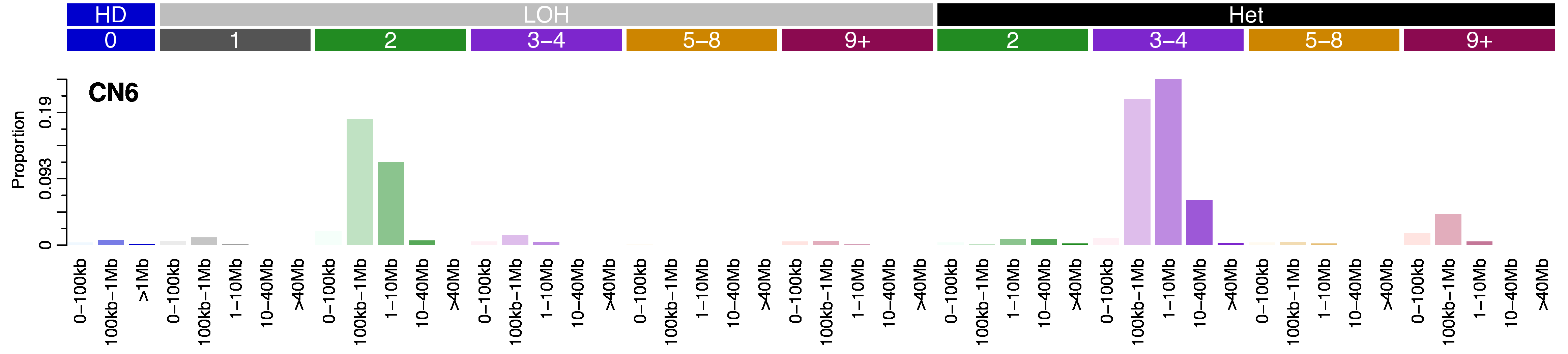
Mutational profile
Copy number signatures are defined by a 48 context copy number classification scheme. The scheme incorporates loss-of-heterozygosity status, total copy number state, and segment length to categorise segments from allele-specific copy number profiles.
The scheme corresponds to three heterozygosity states:
- heterozygous segments with copy number of [A>0, B>0] (numbers reflect the counts for major allele A and minor allele B)
- segments with LOH with copy number of [A>0, B=0]
- segments with homozygous deletions [A=0, B=0]
Segments are further subclassified into 6 classes based on the sum of major and minor allele (total copy number) and chosen for biological relevance:
- total copy number (TCN)=0 (homozygous deletion)
- TCN=1 (deletion leading to LOH)
- TCN=2 (wild type, including copy-neutral LOH)
- TCN=3 or 4 (minor gain)
- TCN=5 to 8 (moderate gain)
- TCN>=9 (high-level amplification).
Each of the heterozygous and LOH total copy number states are then organised into five classes based on the size of their segments: 0 – 100kb, 100kb – 1Mb, 1Mb – 10Mb, 10Mb – 40Mb, and >40Mb (the largest category for homozygous deletions was restricted to >1Mb) in order to capture focal, large scale, and chromosomal-scale copy number changes. This gives a total of 48 copy number context to categorise segments into (3x homozygous deletion categories, 5*5=25x LOH categories, 4*5=20 heterozygous categories).
Proposed aetiology
CN6 is a signature of chromothripsis/amplicon events followed by genome doubling (LOH segments are lost through chromothripsis, and then gained due to genome doubling). Evidence for this comes from associations with local calls of chromothripsis in PCAWG, local calls of chromothripsis before whole genome doubling in PCAWG (see PCAWG consortium, 2020), associations with samples that are once-genome-doubled, and through simulations of chromothripsis followed by genome doubling.
In addition, CN6 associates strongly with the circular amplicon class (ECDNA), and less strongly with the linear and heavily rearranged amplicon classes (see Kim et al., 2020). CN6 is also associated with chromothripsis, deletion and tyfonas rearrangement classes (see Hadi et al., 2020).
Notes
CN6 is associated with a poorer prognosis for patients in a pan-cancer analysis, with a poorer prognosis in just kidney renal cell papillary cancer or just adrenocortical carcinoma, and with a poorer prognosis when taken in combination with all other chromothripsis/amplicon signatures.
CN6 positively associates withPPM1D amplification in breast cancer, and with black ancestry of patients compared to white ancestry.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Steele et al. 2022 Nature | v3.3 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| SNP6 | No | No | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| No | No | - | |
| Proposed aetiology | Mutational process | Support | |
| Chromothripsis | Statistical association | ||
| Experimental validation | Experimental study | Species | |
| - | - | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
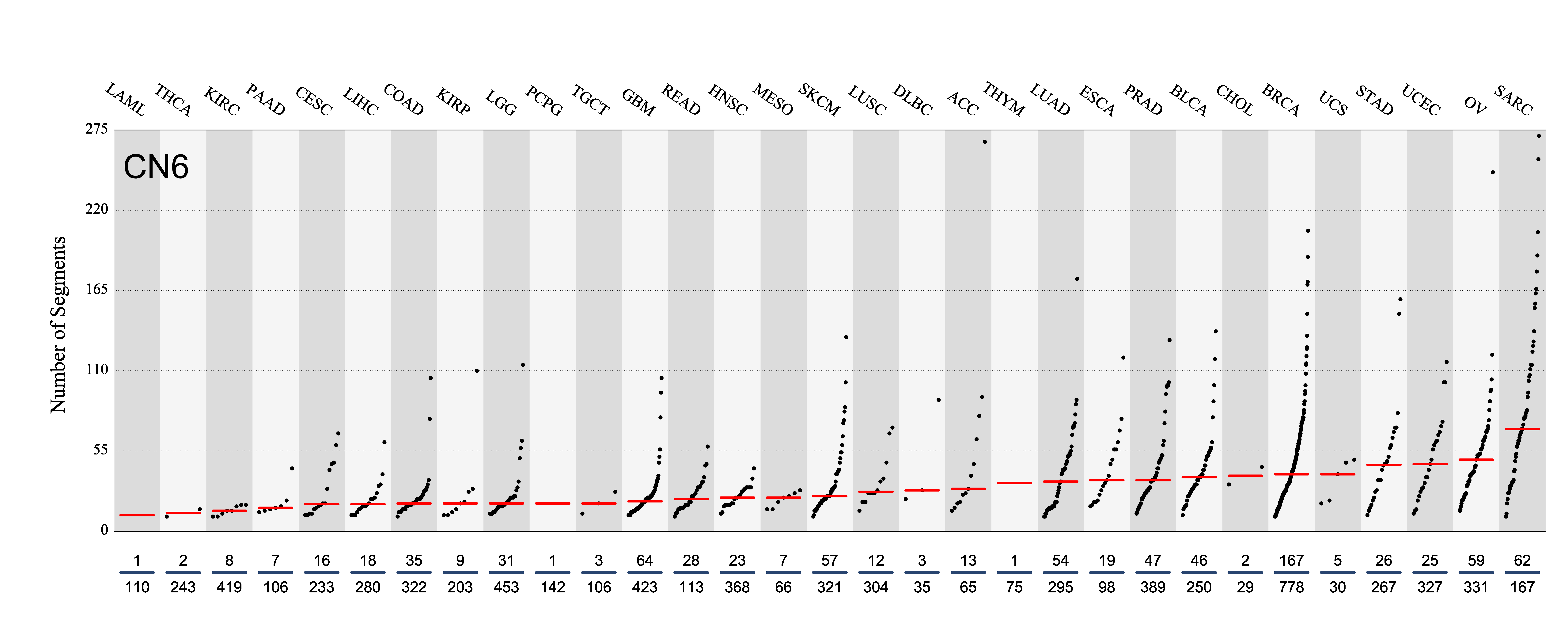
Tissue distribution
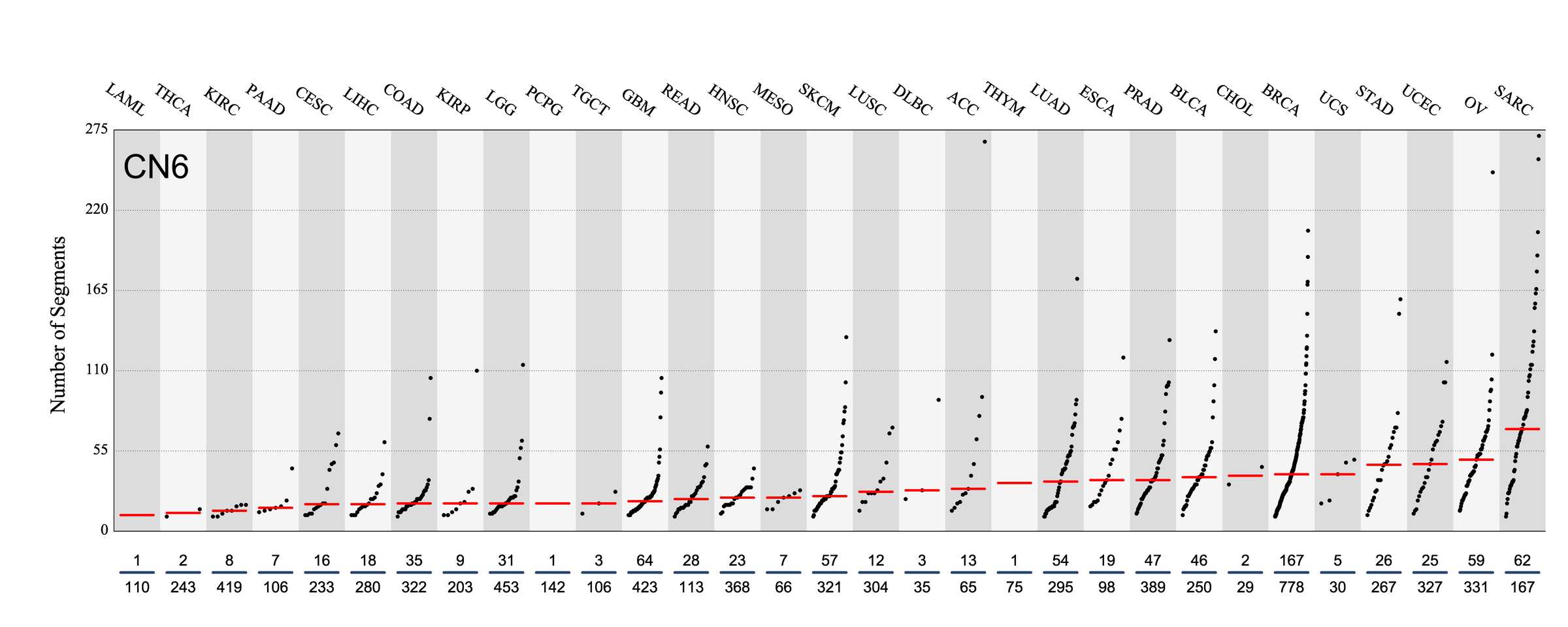
Number of segments attributed to the copy number signature across the cancer types in which it was found. Each dot represents an individual sample and only samples where the signature is found are shown.
The numbers below the dots for each cancer type indicate the number of high confidence tumours in which at least 10 mutations were attributed to the signature (above the blue horizontal line) and the total number of high confidence tumours analysed (below the blue horizontal line). Only high confidence data are displayed: samples with reconstruction accuracy >0.90. Note that due to the nature of copy number data, a sample with e.g. 22 counts of CN1 (a majority diploid unaltered signature, with large segment sizes) may span the whole genome, while a sample with e.g. 70 counts of CN8 (a highly segmented chromothripsis amplification signature, with small segment sizes) may span a small region of the genome.