Mutational Signatures Version 2 (March 2015)
Signatures of Mutational Processes in Human Cancer
Signatures of Mutational Processes in Human Cancer
This page shows version 2 of the Mutational Signatures. You can see the updated and expanded set of signatures in version 3.2 (March 2021).
Somatic mutations are present in all cells of the human body and occur throughout life. They are the consequence of multiple mutational processes, including the intrinsic slight infidelity of the DNA replication machinery, exogenous or endogenous mutagen exposures, enzymatic modification of DNA and defective DNA repair. Different mutational processes generate unique combinations of mutation types, termed “Mutational Signatures”.
In the past few years, large-scale analyses have revealed many mutational signatures across the spectrum of human cancer types [Nik-Zainal S. et al., Cell (2012); Alexandrov L.B. et al., Cell Reports (2013); Alexandrov L.B. et al., Nature (2013); Helleday T. et al., Nat Rev Genet (2014); Alexandrov L.B. and Stratton M.R., Curr Opin Genet Dev (2014)]. However, as the number of mutational signatures grows the need for a curated census of signatures has become apparent. Here, we deliver such a resource by providing the profiles of, and additional information about, known mutational signatures.
The current set of mutational signatures is based on an analysis of 10,952 exomes and 1,048 whole-genomes across 40 distinct types of human cancer. These analyses are based on curated data that were generated by The Cancer Genome Atlas (TCGA), the International Cancer Genome Consortium (ICGC), and a large set of freely available somatic mutations published in peer-reviewed journals. Complete details about the data sources will be provided in future releases of COSMIC.
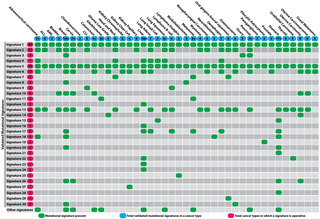
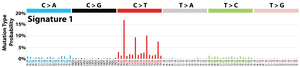
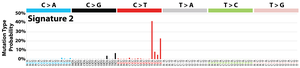
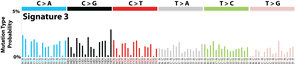
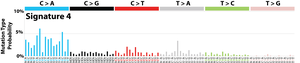
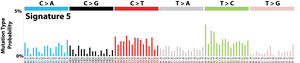
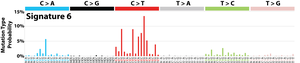
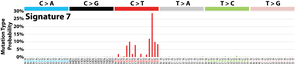
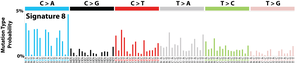
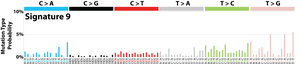
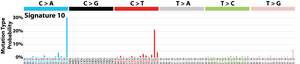
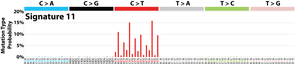
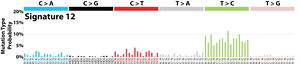
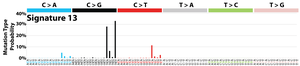
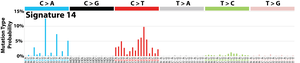
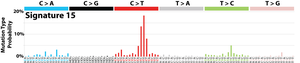
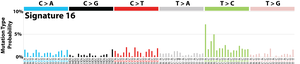
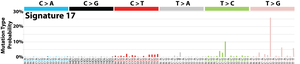
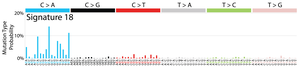
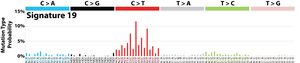
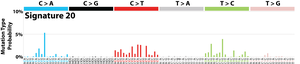
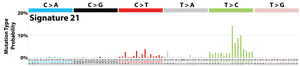
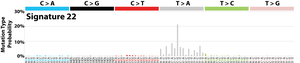
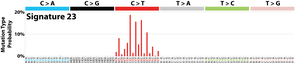
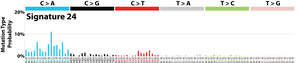
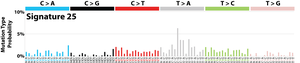
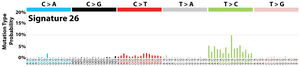
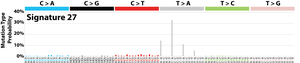
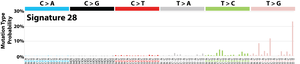
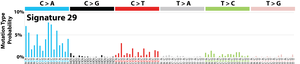
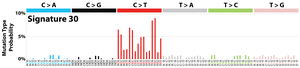
The profile of each signature is displayed using the six substitution subtypes: C>A, C>G, C>T, T>A, T>C, and T>G (all substitutions are referred to by the pyrimidine of the mutated Watson–Crick base pair). Further, each of the substitutions is examined by incorporating information on the bases immediately 5’ and 3’ to each mutated base generating 96 possible mutation types (6 types of substitution ∗ 4 types of 5’ base ∗ 4 types of 3’ base). Mutational signatures are displayed and reported based on the observed trinucleotide frequency of the human genome, i.e., representing the relative proportions of mutations generated by each signature based on the actual trinucleotide frequencies of the reference human genome version GRCh37. Note that only validated mutational signatures have been included in the curated census of mutational signatures.
Additional information is provided for each signature, including the cancer types in which the signature has been found, proposed aetiology for the mutational processes underlying the signature, other mutational features that are associated with each signature and information that may be relevant for better understanding of a particular mutational signature.
All the data used for deriving the 30 mutational signatures in COSMIC are publicly and freely available. Please note that most of these data were generated as part of studies performed by others. Information about the data sources containing these freely available data is provided in the supplementary information of “Clock-like mutational processes in human somatic cells” Nature Genetics, 2015, 47, 1402-1407 , the supplementary information of “Landscape of somatic mutations in 560 breast cancer whole-genome sequences” Nature, 534, 47–54 , the supplementary information of “Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups” Nature Communications, 4, Article number: 2873 , and the COSMIC Cell Lines Project. This information can be used to download any sample from its respective data source. An updated set of mutational matrices allowing reproduction of the COSMIC signatures found in primary cancers can be downloaded from here . The cancer cell line data, from which signature 25 was derived, can be retrieved from the COSMIC Cell Lines Project.
The set of signatures has been updated in version 3.2 (March 2021), incorporating additional mutation types (indels and doublet base substitutions) and cancer samples.
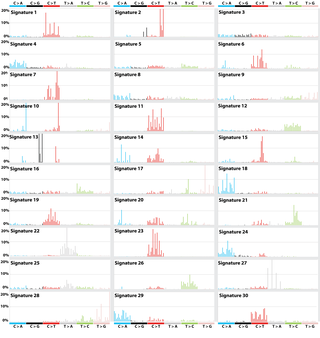
Patterns of mutational signatures [Download signatures]