Mutational Signatures (v3.6 - May 2026)
SV1 · GRCh38 · COSMIC v104
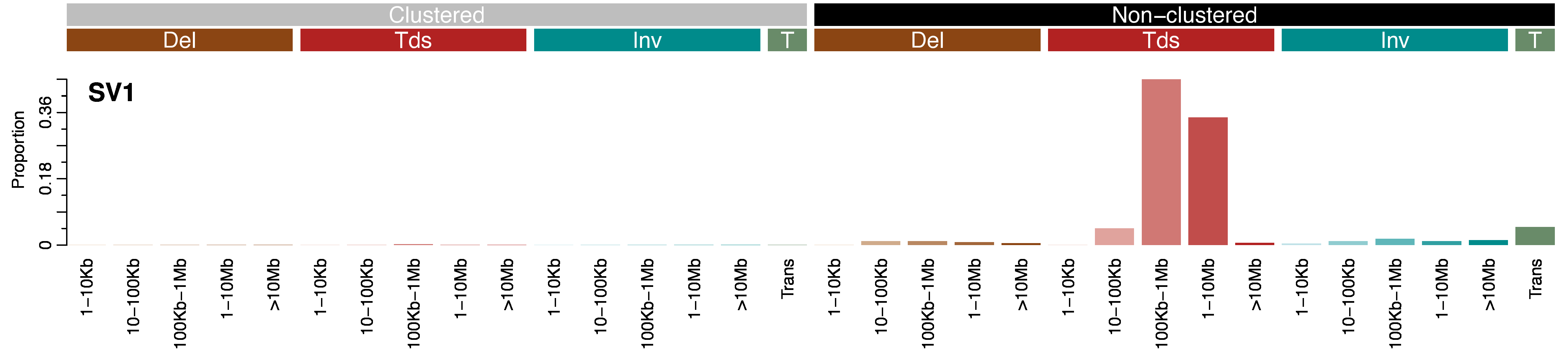
Mutational profile
Overview of the SV32 classification schema. The classification schema bins all SVs, apart from translocations, according to the size of the event in base pairs: 0–10kb, 10kb–100kb, 100kb–1Mb, 1Mb–10Mb, and >10Mb. Translocations, which may involve more than one chromosome, are not binned by size because they can be either balanced (where there is no net loss of genetic material on the chromosomes involved and thus the size can be described by one number) or unbalanced (where there is a net loss or gain of genetic material on the chromosomes involved and thus the sizes of the segments cannot be described by just one number). Note that whether a translocation is balanced or unbalanced is not considered in this classification schema. The different types of SVs are then further divided into clustered and non-clustered events to account for the non-random distribution of these events along the genome. Clustered events are defined as events that occur closer to each other on a chromosome than purely expected by chance.
Proposed aetiology
Unknown.
Comments
SV1 exhibits long tandem duplications ranging from 100Kb to 10Mb. This signature is predominantly observed in breast, ovarian, and uterine cancers. The signature has been previously identified independently in Nik-Zainal et al. 2016 (RS1).
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Nik-Zainal et al. 2016 Nature | v3.4 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | Yes | - | |
| Proposed aetiology | Mutational process | Support | |
| Unknown | Unknown | ||
| Experimental validation | Experimental study | Species | |
| - | - | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Found in bladder, breast, CNS, hepatopancreatobiliary, lung, ovary, prostate, skin, upper gastrointestinal, and uterine cancers.