Mutational Signatures (v3.6 - May 2026)
RNA-SBS2 · GRCh37 · COSMIC v104
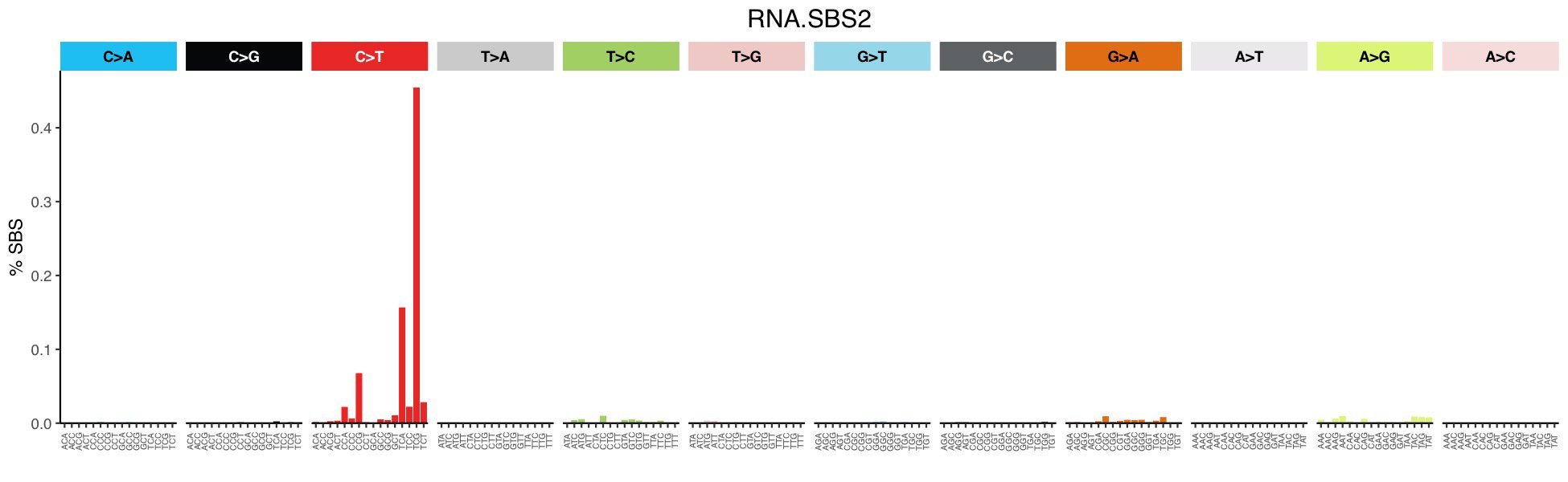
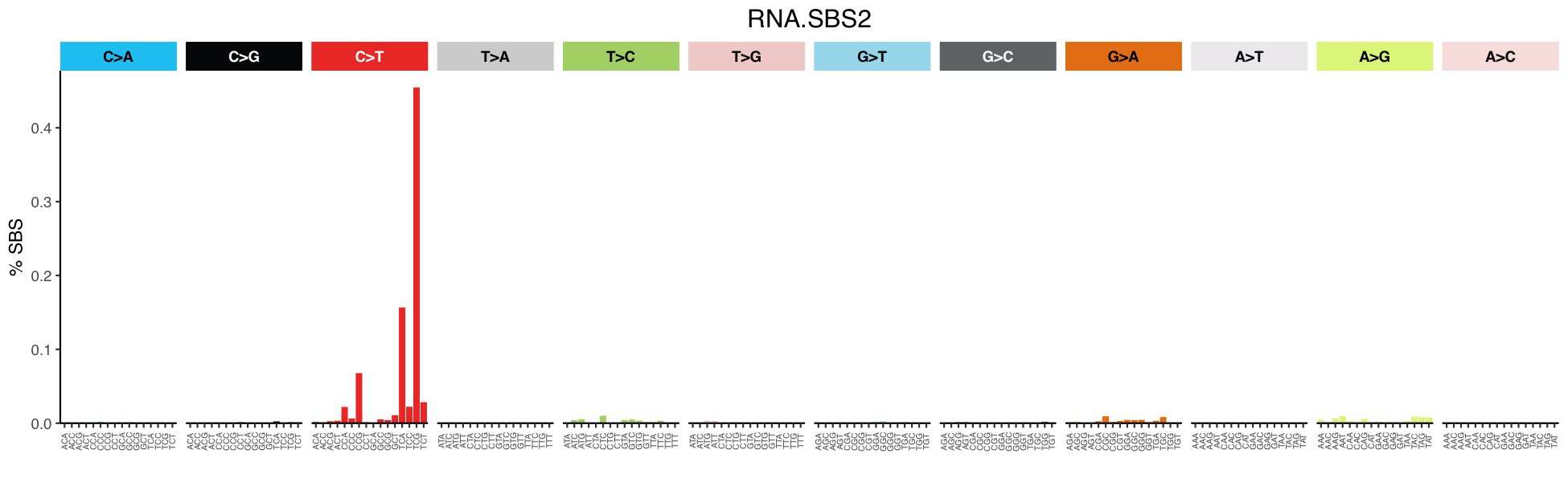
Mutational profile
Mutational profile using 192-channels. This classification is defined by using the 192-channel for the tri-nucleotide context of every possible point nucleotide change on an RNA molecule. Because RNA molecules are single-stranded, we can infer the exact nucleotide change, thus resulting in the 192 channel, unlike DNA single base substitutions, where the strandedness of the nucleotide change cannot always be inferred.
Proposed aetiology
Consisted mainly of C>T transitions at TpC sites relating to RNA editing activity of APOBEC3A.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Martinez-Ruiz et al. 2023 Nature | v3.4 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| RNA-seq/WES | No | No | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | No | - | |
| Proposed aetiology | Mutational process | Support | |
| RNA editing activity of APOBEC3A | Statistical association | ||
| Experimental validation | Experimental study | Species | |
| Fixman et al. 2024 JMB | Human | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Found in non-small cell lung cancer (NSCLC) tumours from patients in the TRACERx study.