Mutational Signatures (v3.6 - May 2026)
ID4 · GRCh37 · COSMIC v104
Mutational profile
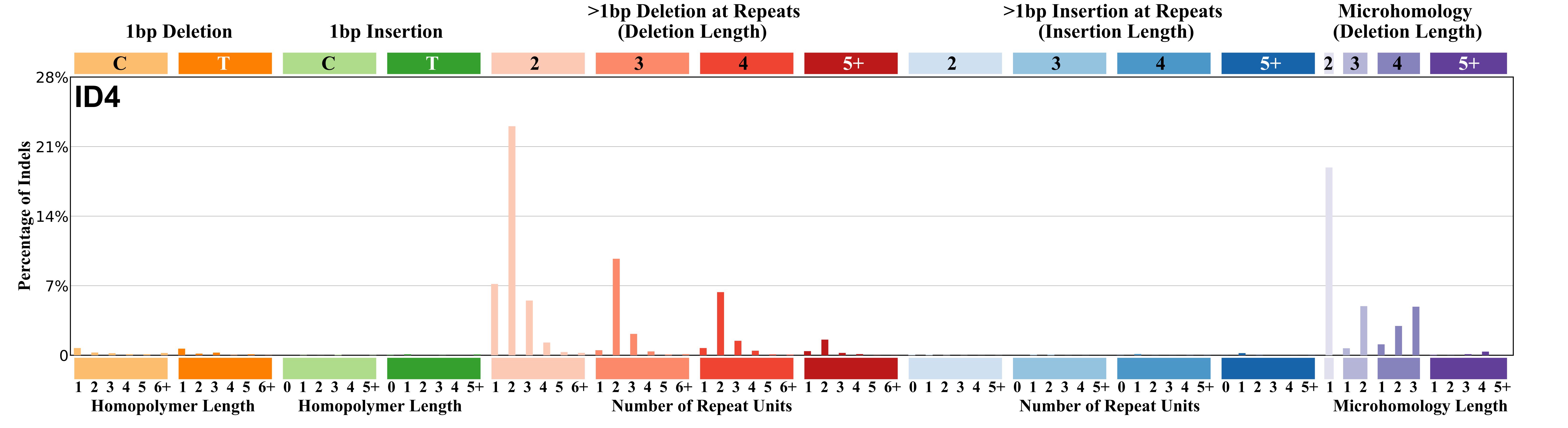
The height of each mutational profile bar represents the proportion of one ID mutation type among all ID mutation types in the signature. Although there is no single intuitive and naturally constrained set of ID mutation types (as there arguably are for SBSs and DBSs), an 83 subclass categorisation of ID mutations was designed.
The 83 ID classification incorporates the prior knowledge that IDs commonly have sizes of 1-10 bps, that both insertions and deletions exist, that IDs of C and T occur at different rates, that IDs preferentially occur at repetitive elements, that the length of the repeat unit may influence the likelihood of an ID occurring, that the number of repeat units in a repeat stretch may influence the likelihood of an ID occurring, that IDs are also fostered in some instances by overlapping sequence microhomologies at the ID boundaries and that different mutational processes may, in principle, be differently influenced by these features. We therefore designed an 83 subclass categorisation of IDs that allows some exploration of all the above possibilities, while constraining the number of categories in order to accommodate the relatively small numbers of IDs (compared to substitutions) found in most genomes. This classification categorises IDs of lengths from 1bp to >5bp, for 1bp IDs classifies them as T or C and the number of single base repeats they occur in from 0 to >5, categorises lengths of non-single base repeat units from 2bp to >5bp and the number of repeats from 1 to >5 and size of microhomology from 0bp to >5bp. We recognise that different classifications of IDs may be preferred by others. The ID mutation types are enumerated in the following Excel document.
Proposed aetiology
Unknown.
Comments
ID4 shows large numbers of mutations in a subset of samples which do not obviously have the mutational features of defective DNA mismatch repair.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Alexandrov et al. 2020 Nature | v3 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WES & WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | Yes | - | |
| Proposed aetiology | Mutational process | Support | |
| Unknown | Unknown | ||
| Experimental validation | Experimental study | Species | |
| - | - | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
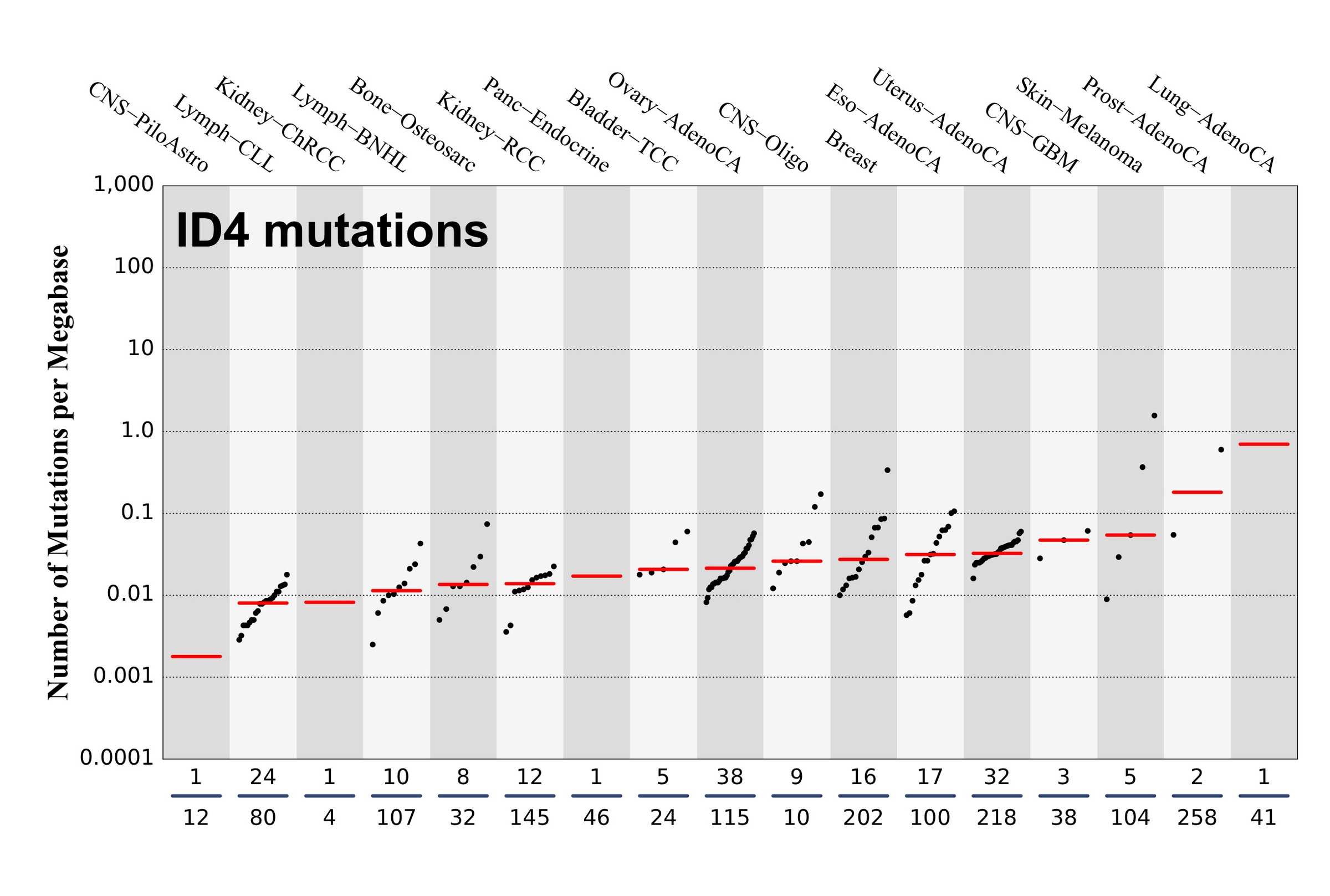
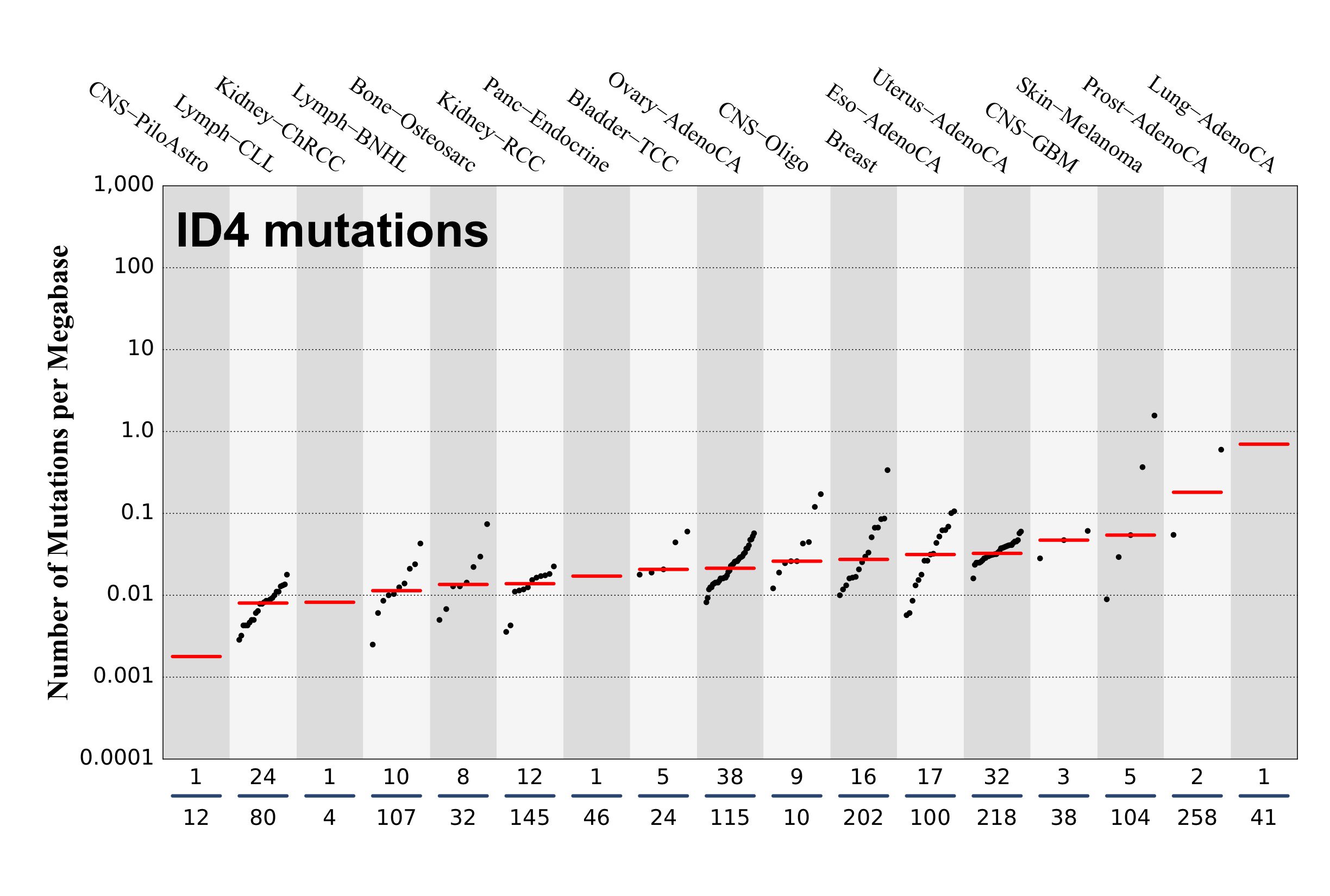
Numbers of mutations per megabase attributed to the mutational signature across the cancer types in which the signature was found. Each dot represents an individual sample and only samples where the signature is found are shown. The number of mutations per megabase was calculated by assuming that an average whole-exome has 30 Mb with sufficient coverage, whereas an average whole-genome has 2,800 Mb with sufficient coverage.
The numbers below the dots for each cancer type indicate the number of high confidence tumours in which the signature was attributed (above the blue horizontal line) and the total number of high confidence tumours analysed (below the blue horizontal line). Only high confidence data are displayed: samples with reconstruction accuracy >0.90.
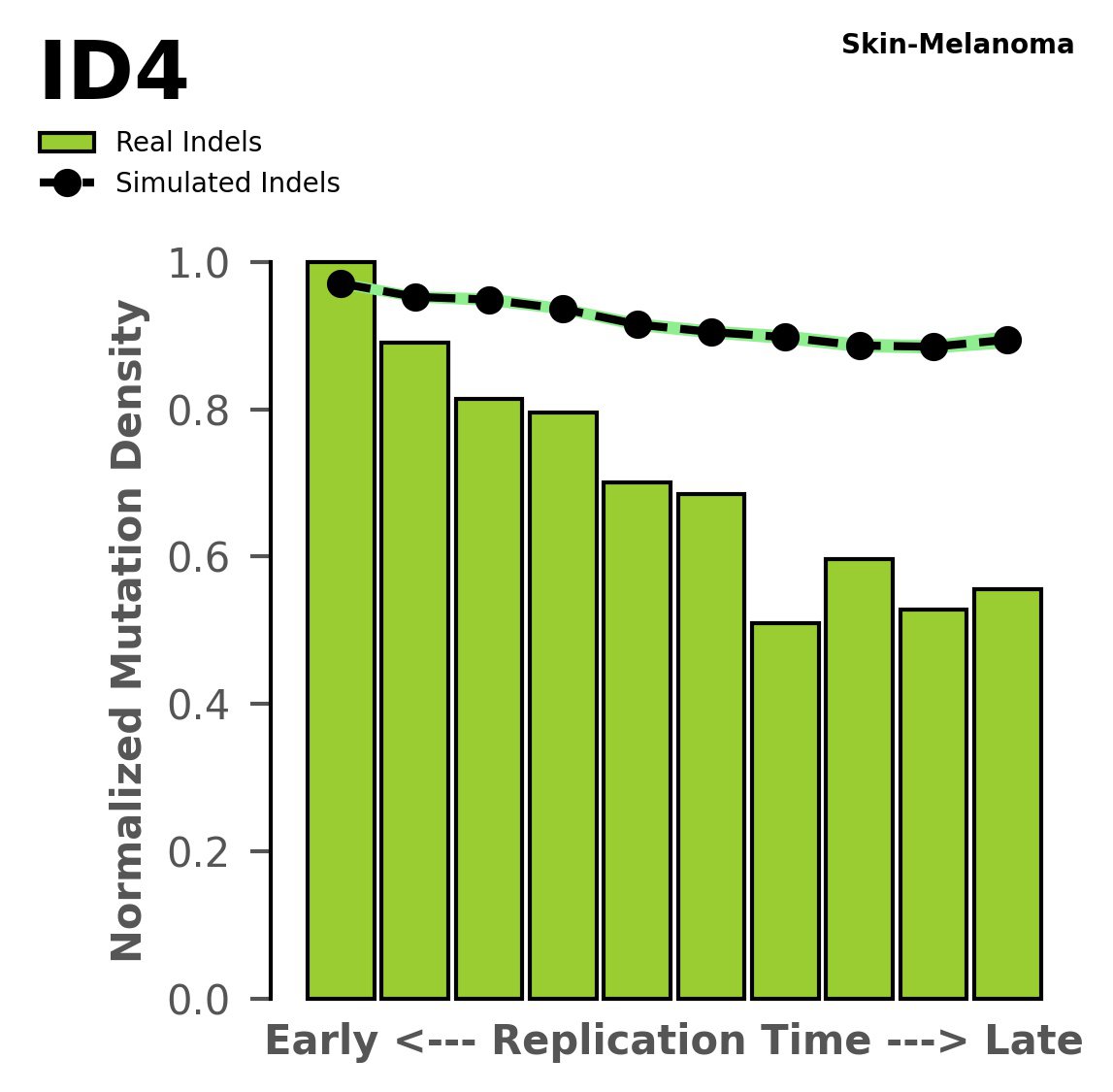
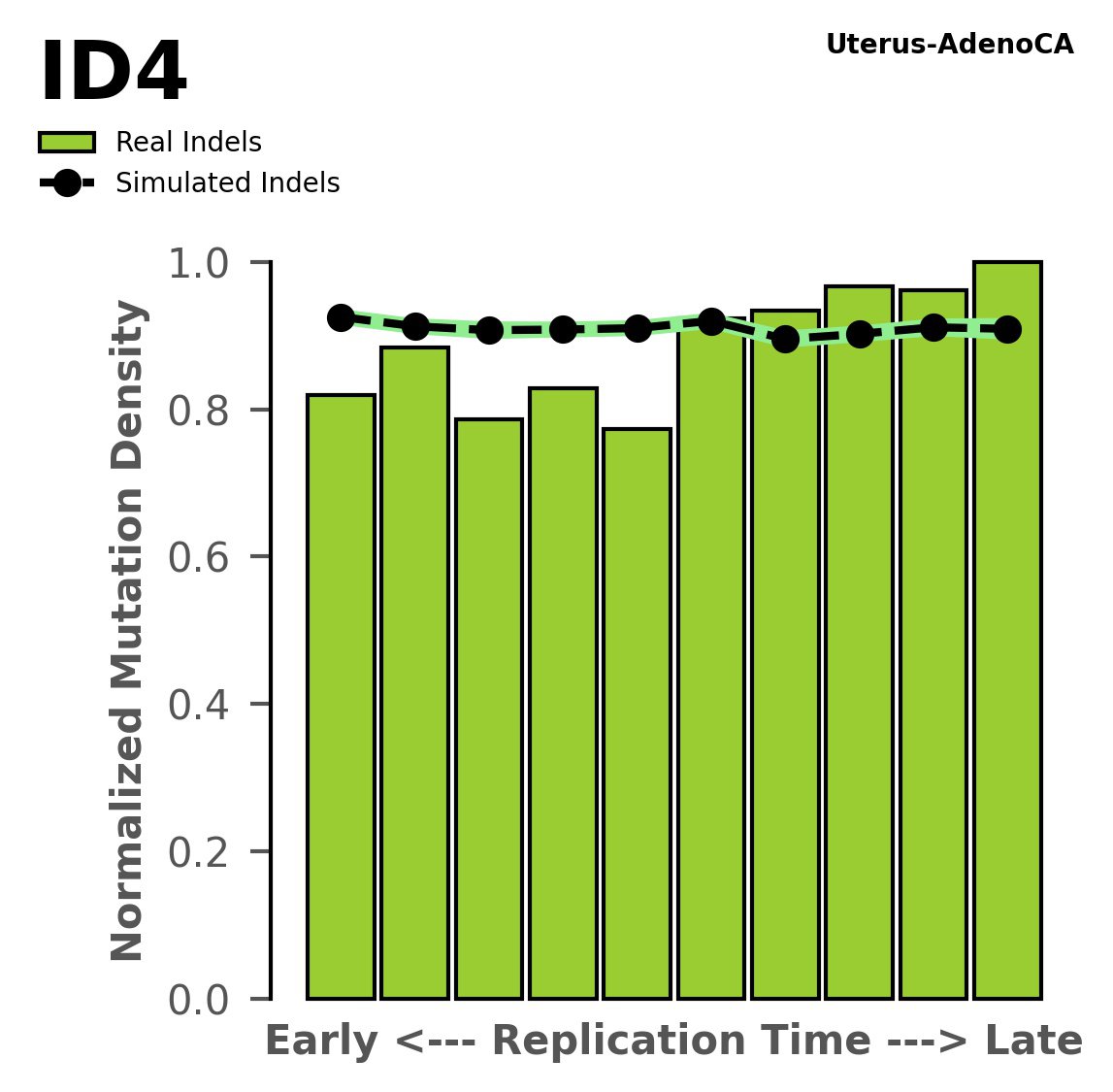
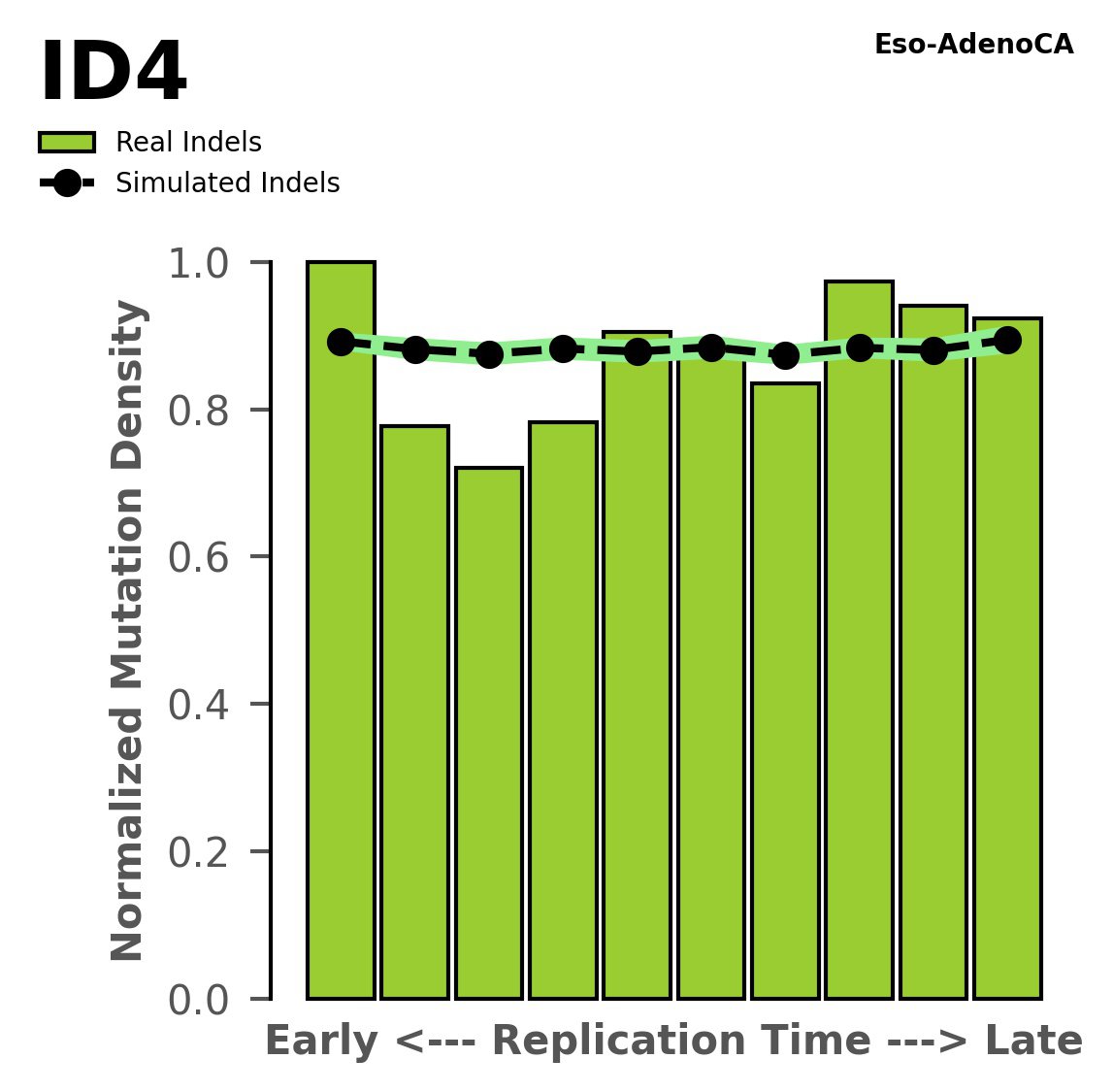
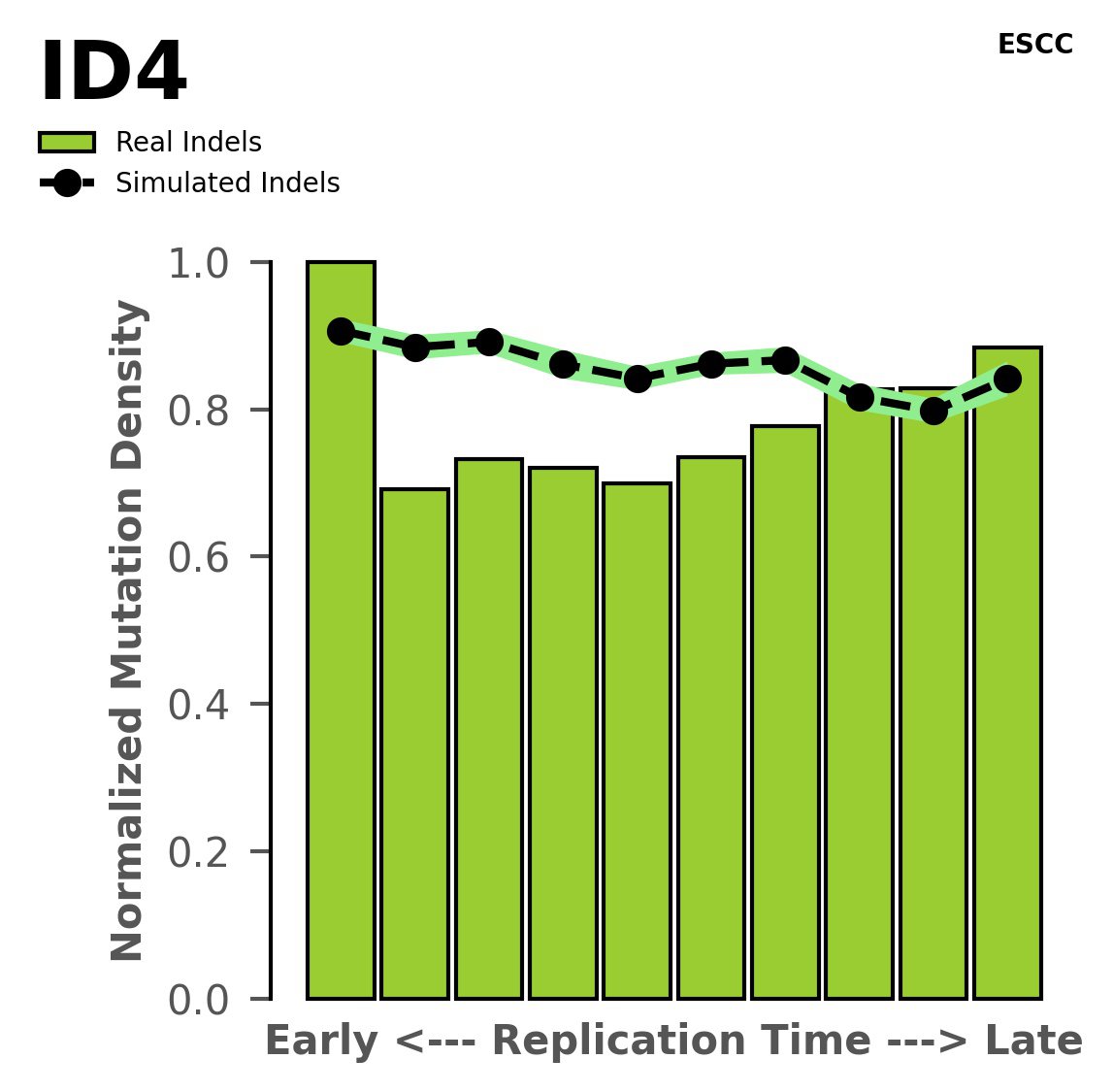
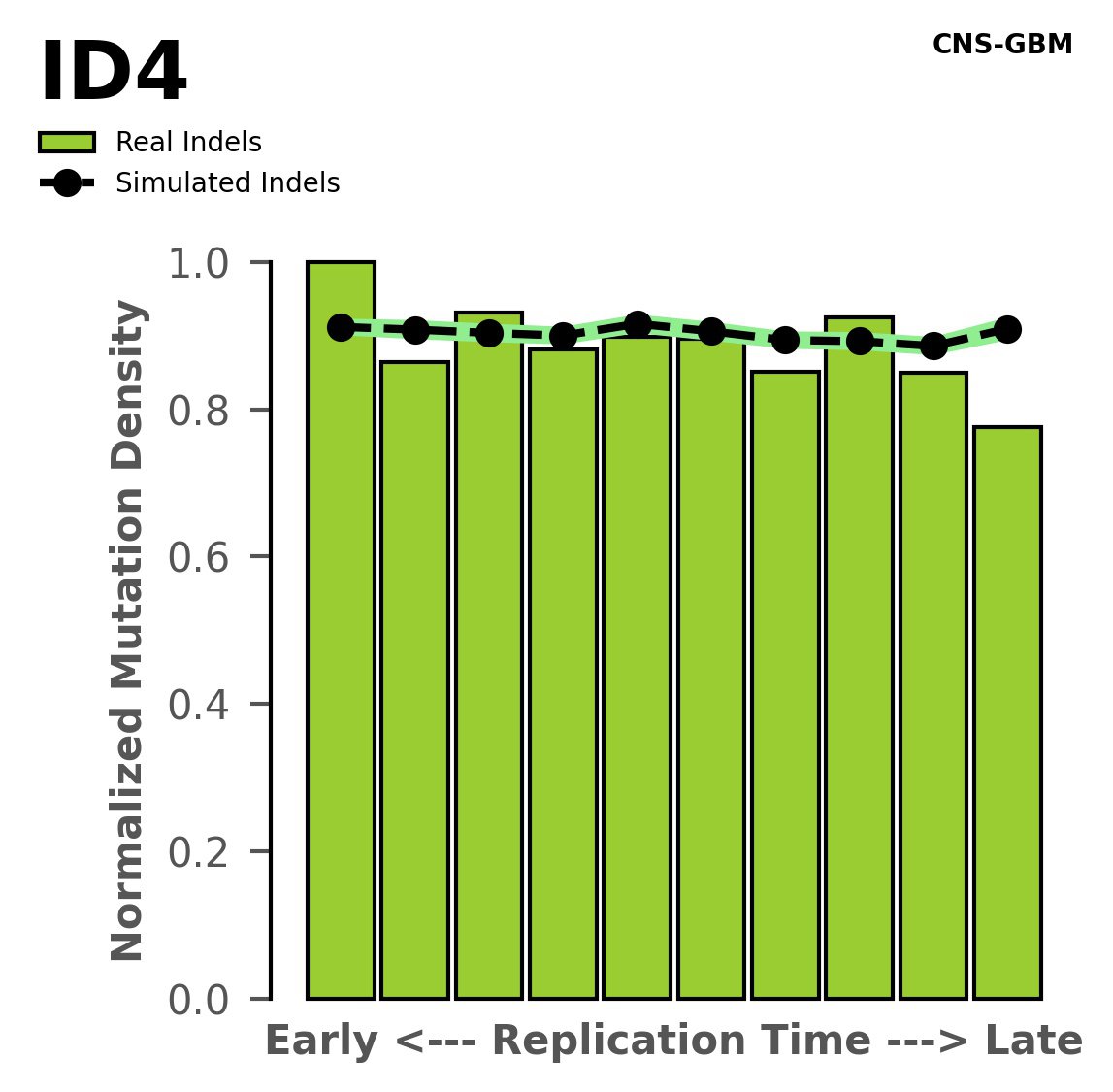
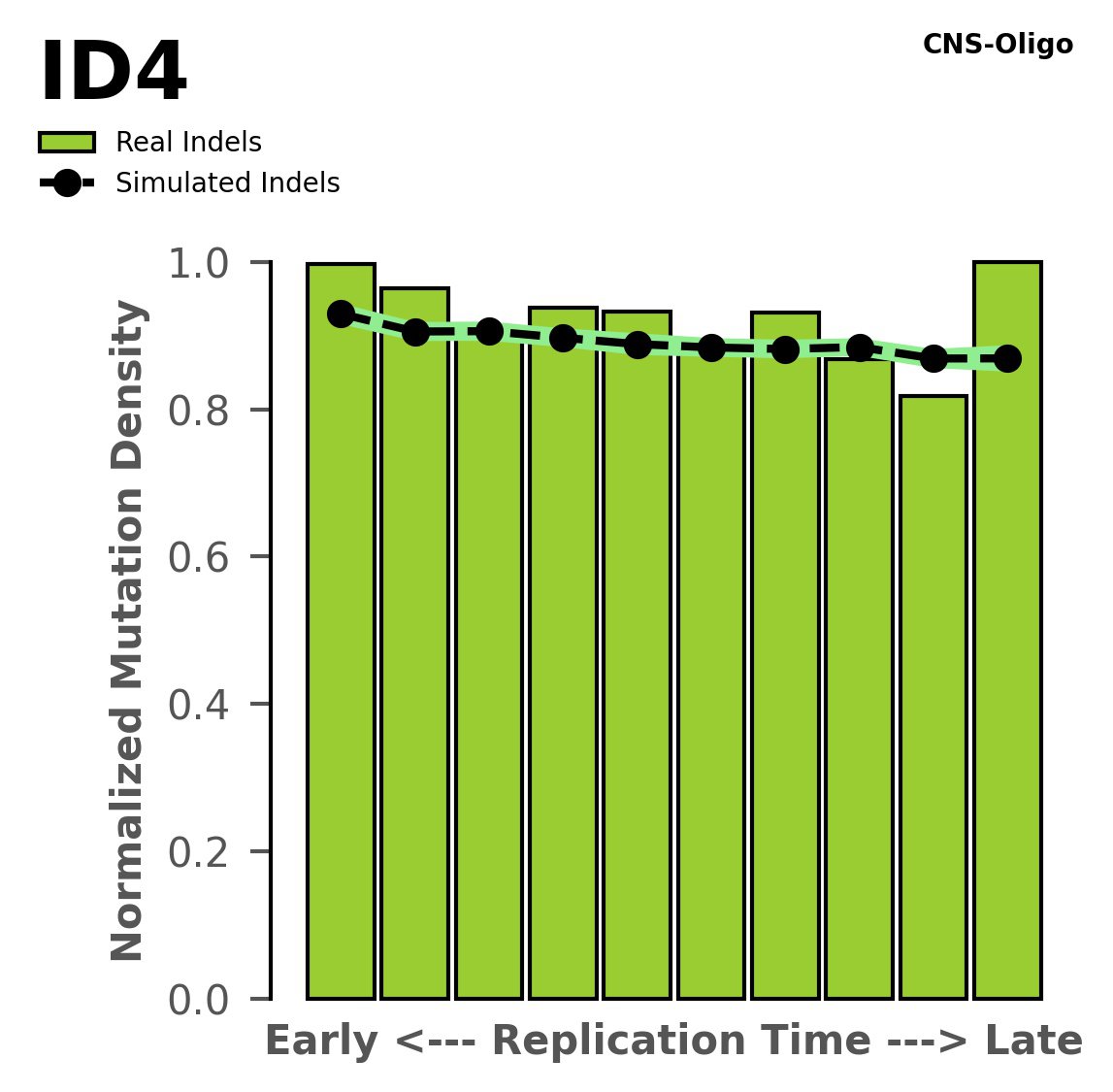
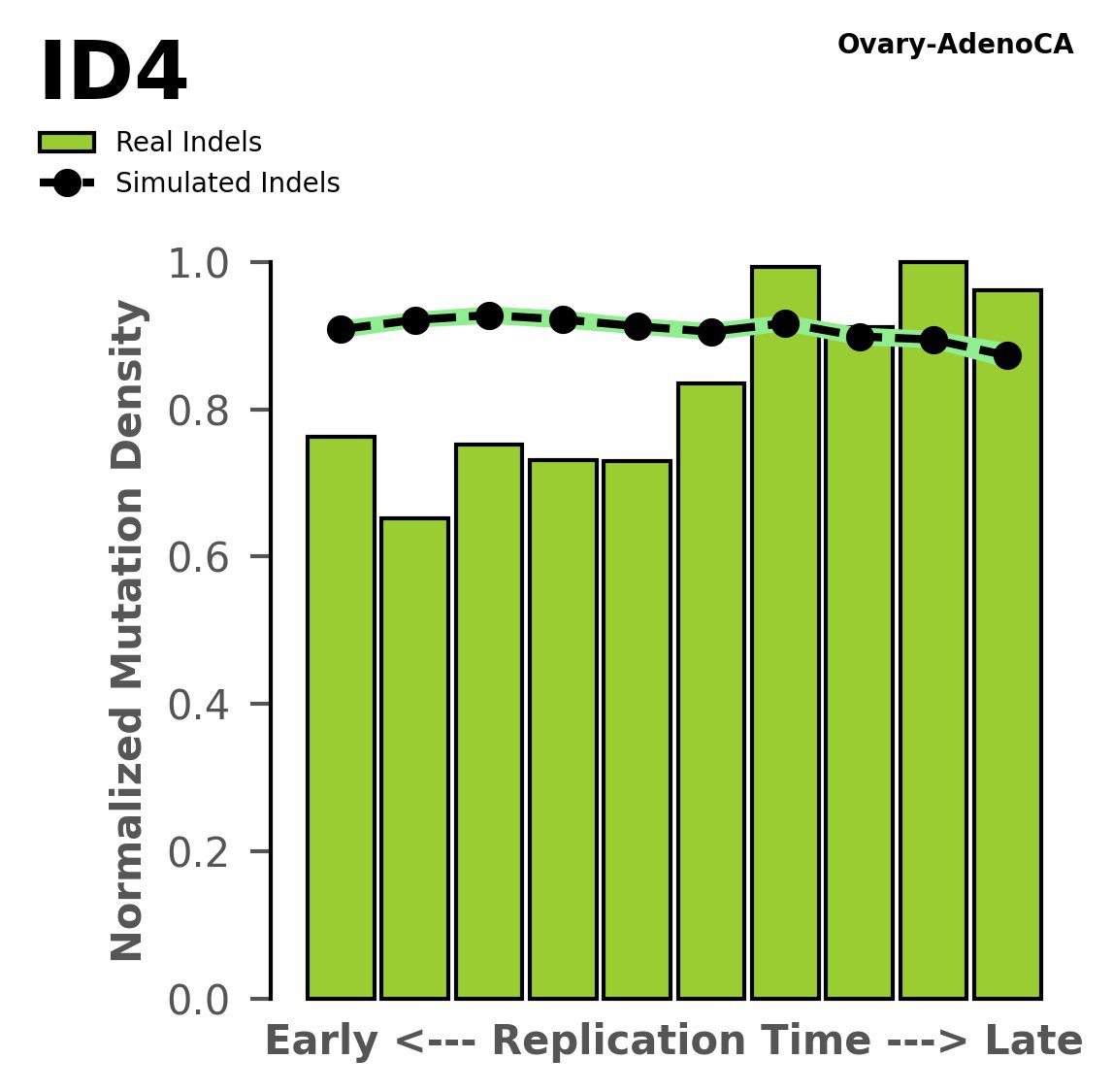
Replication timing
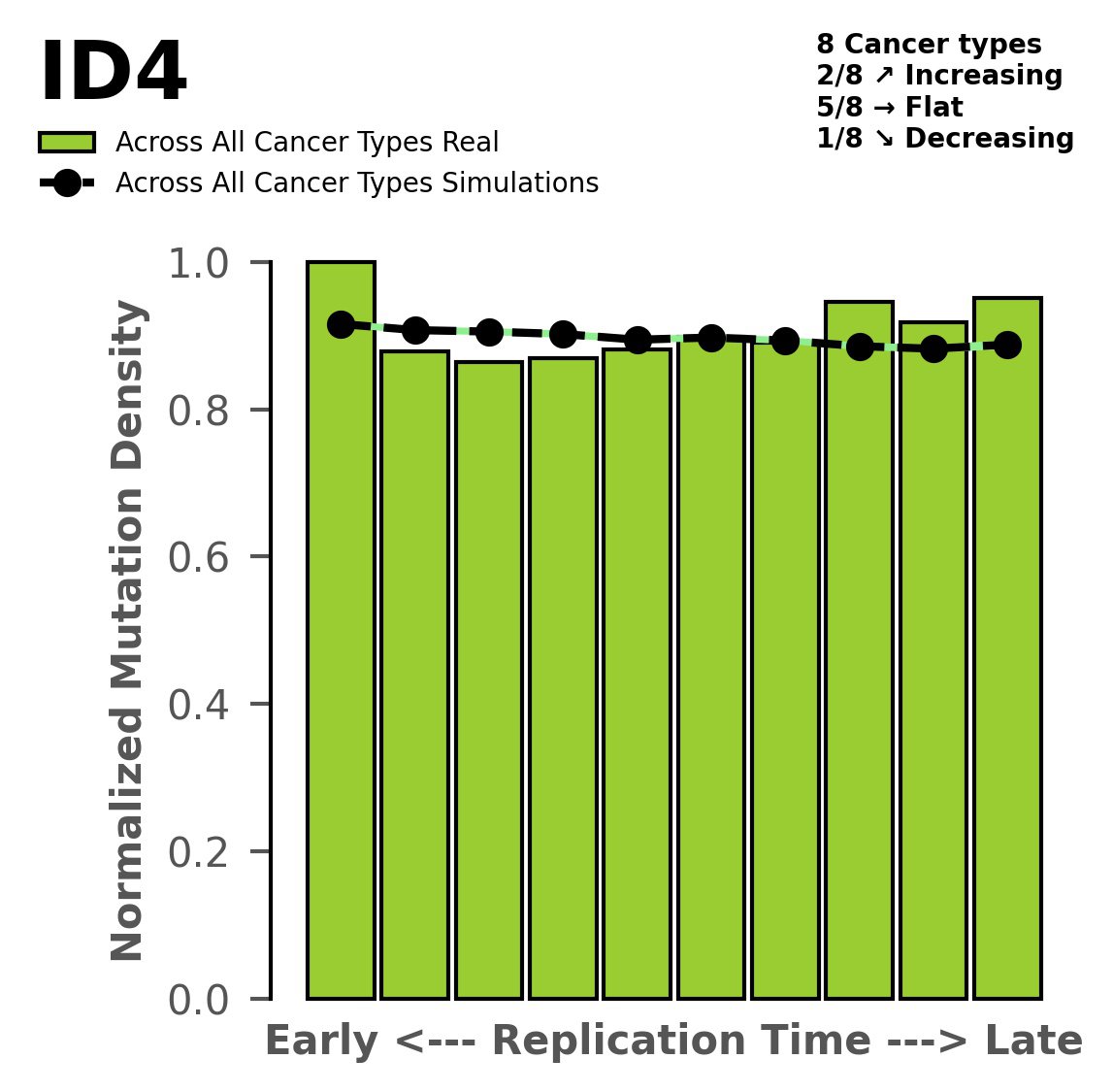
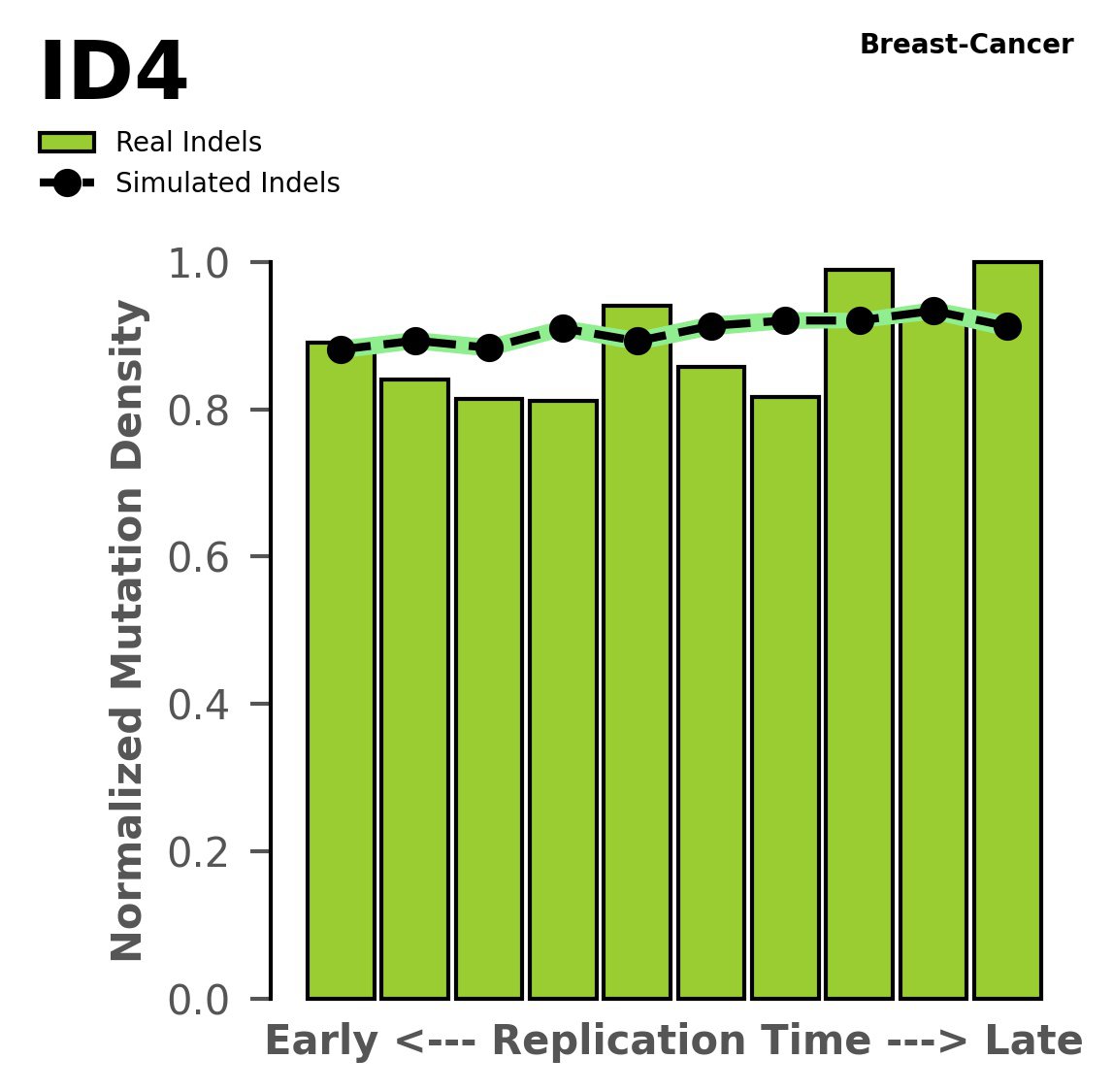
Normalised mutational densities from early to late replicating regions in the human genome are shown with respect to real somatic mutations and simulated mutations. The dashed line reflects the behaviour of simulated mutations, whereas the bars represent the behaviour for real somatic mutations.
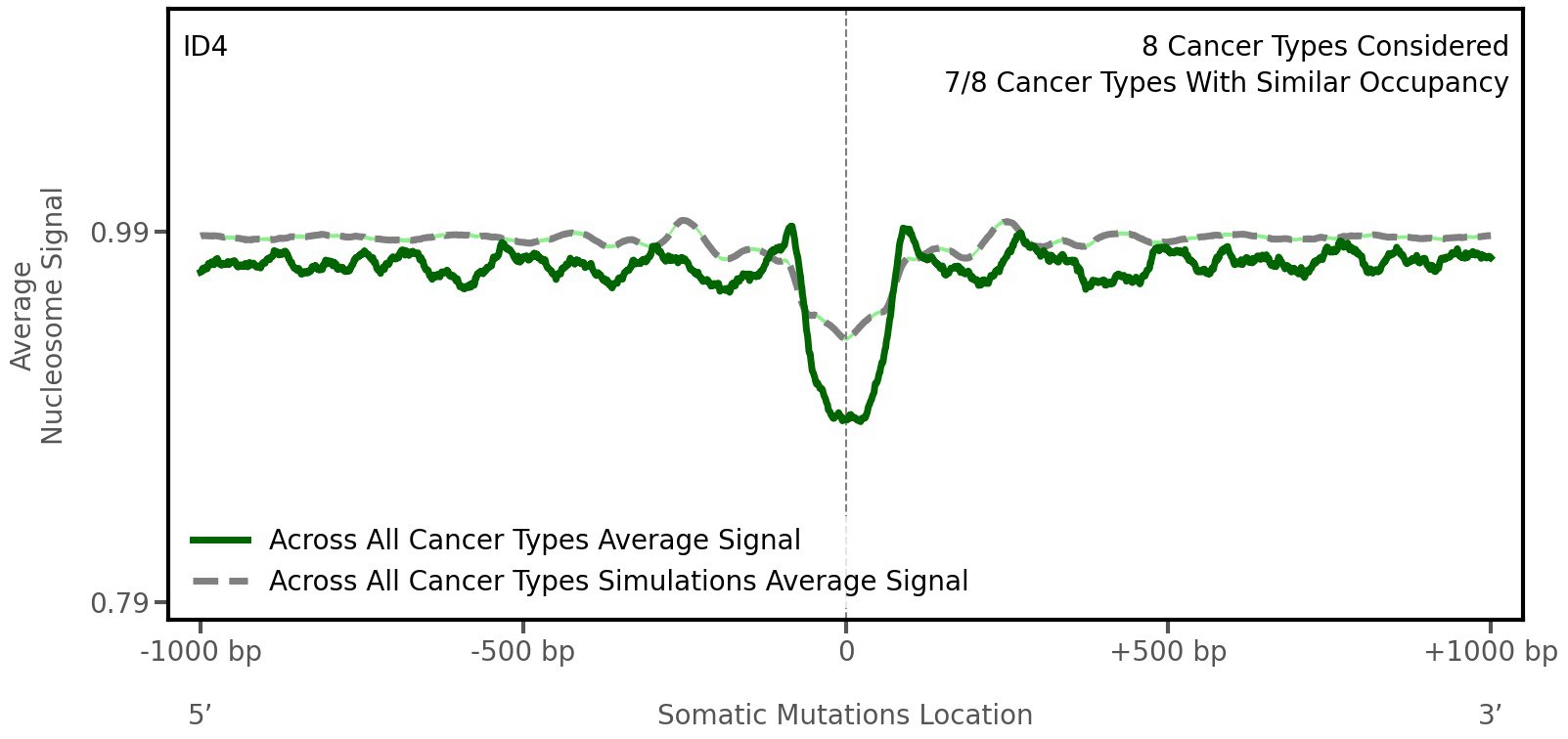
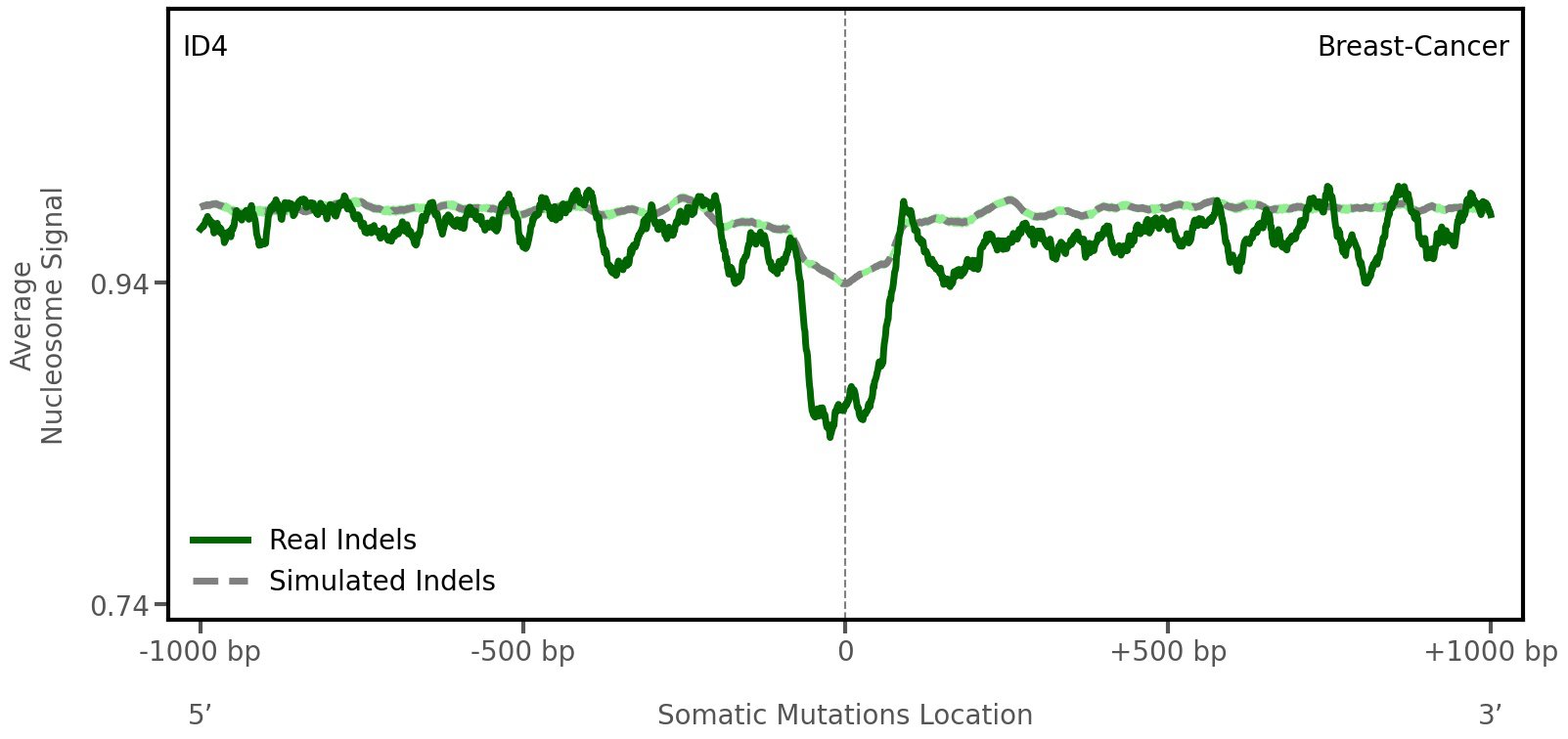
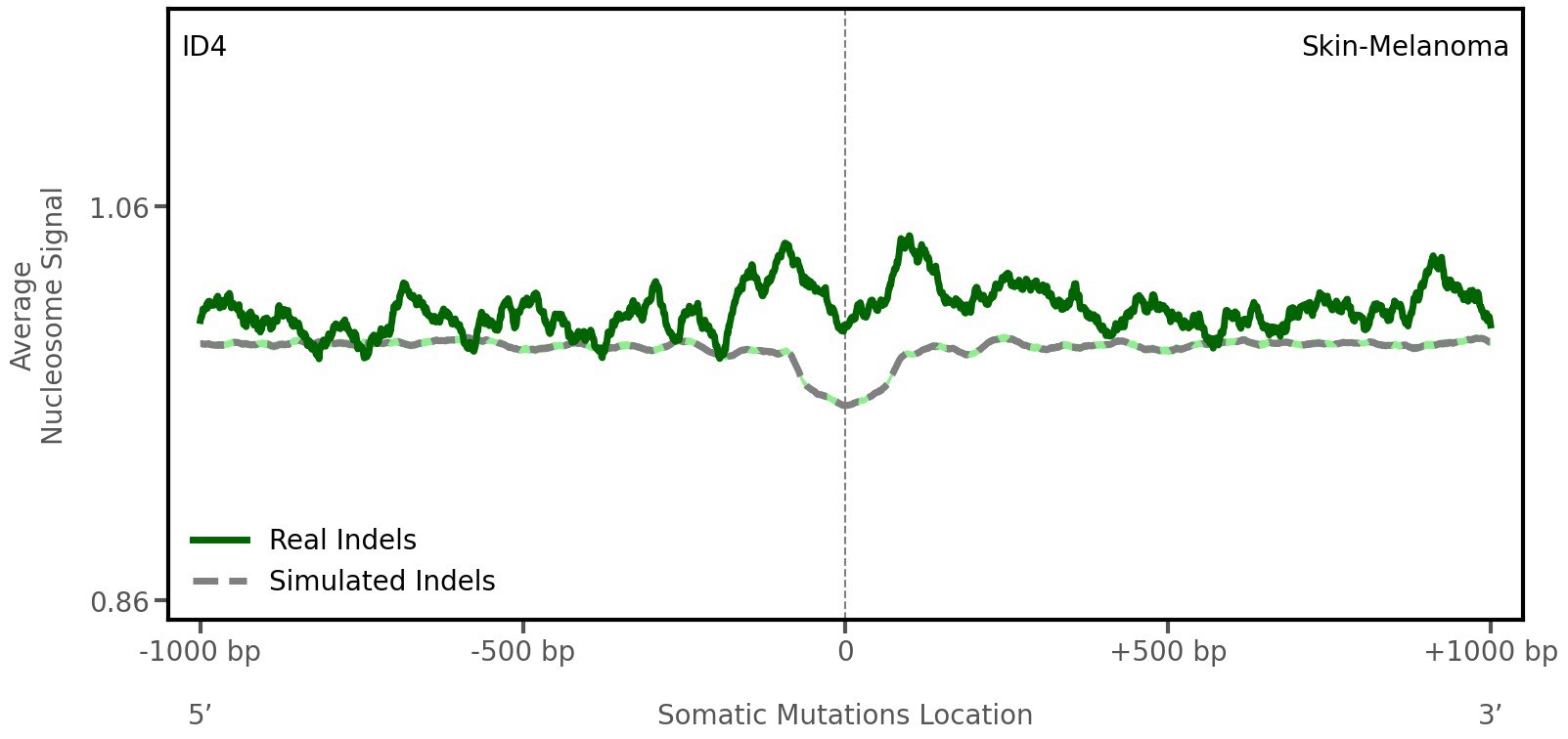
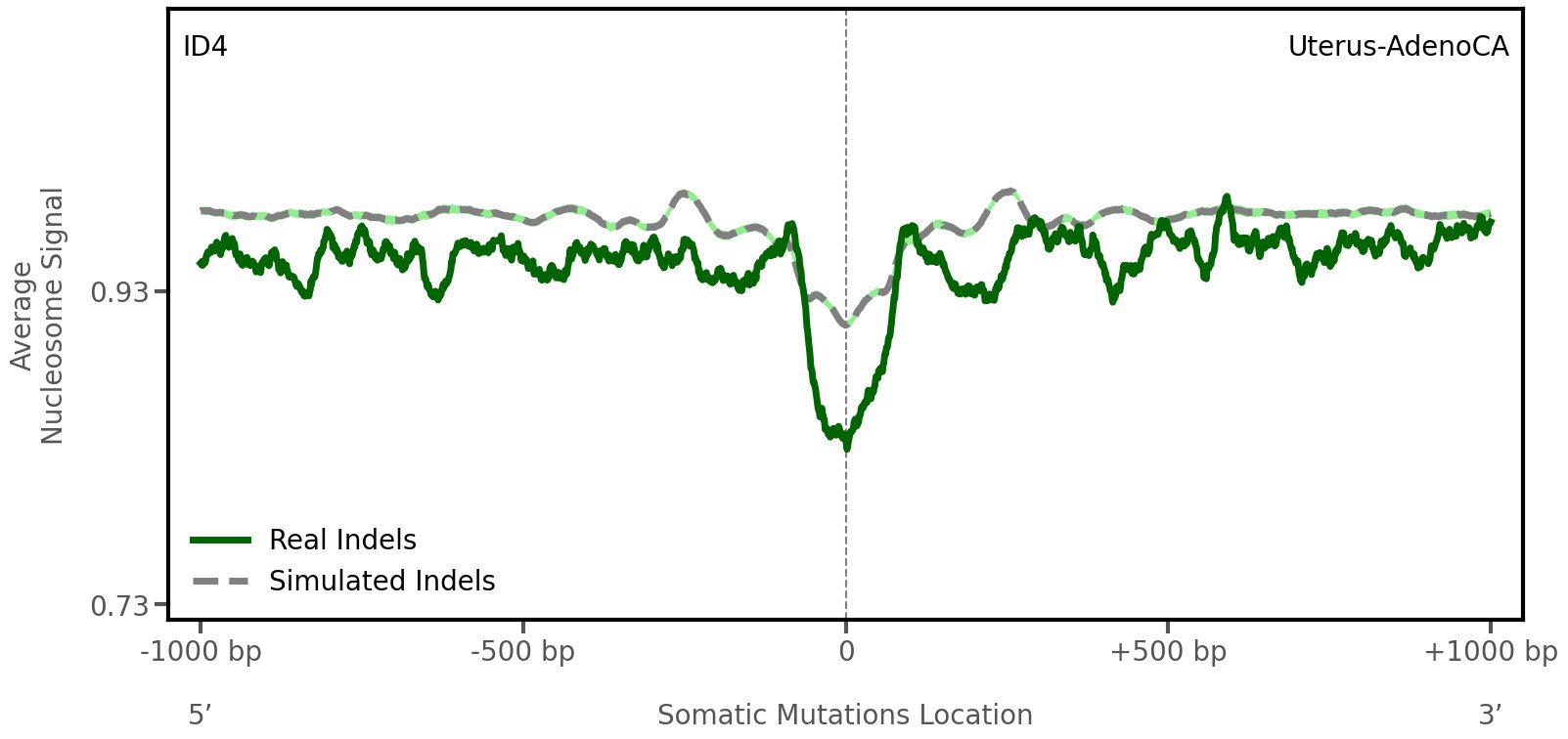
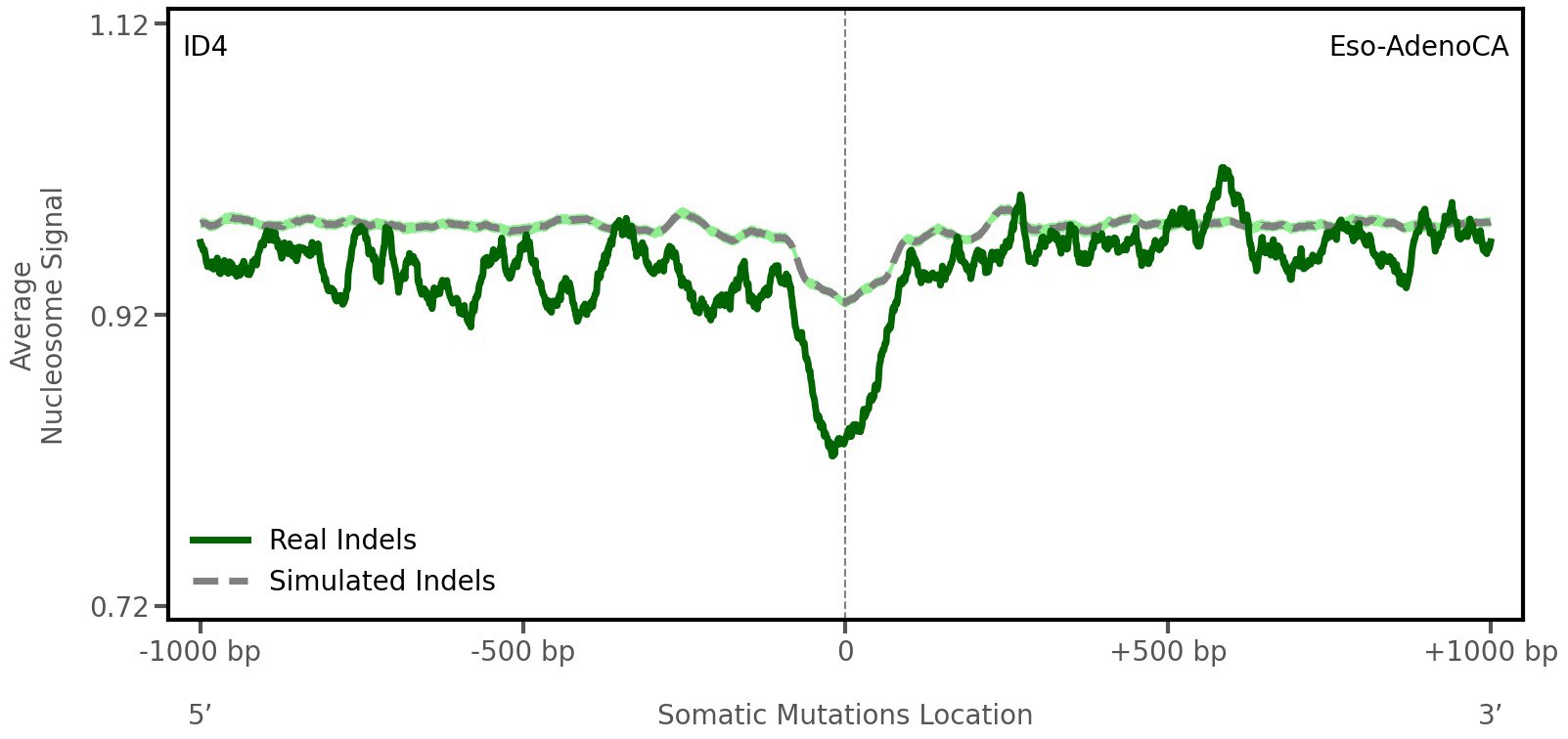
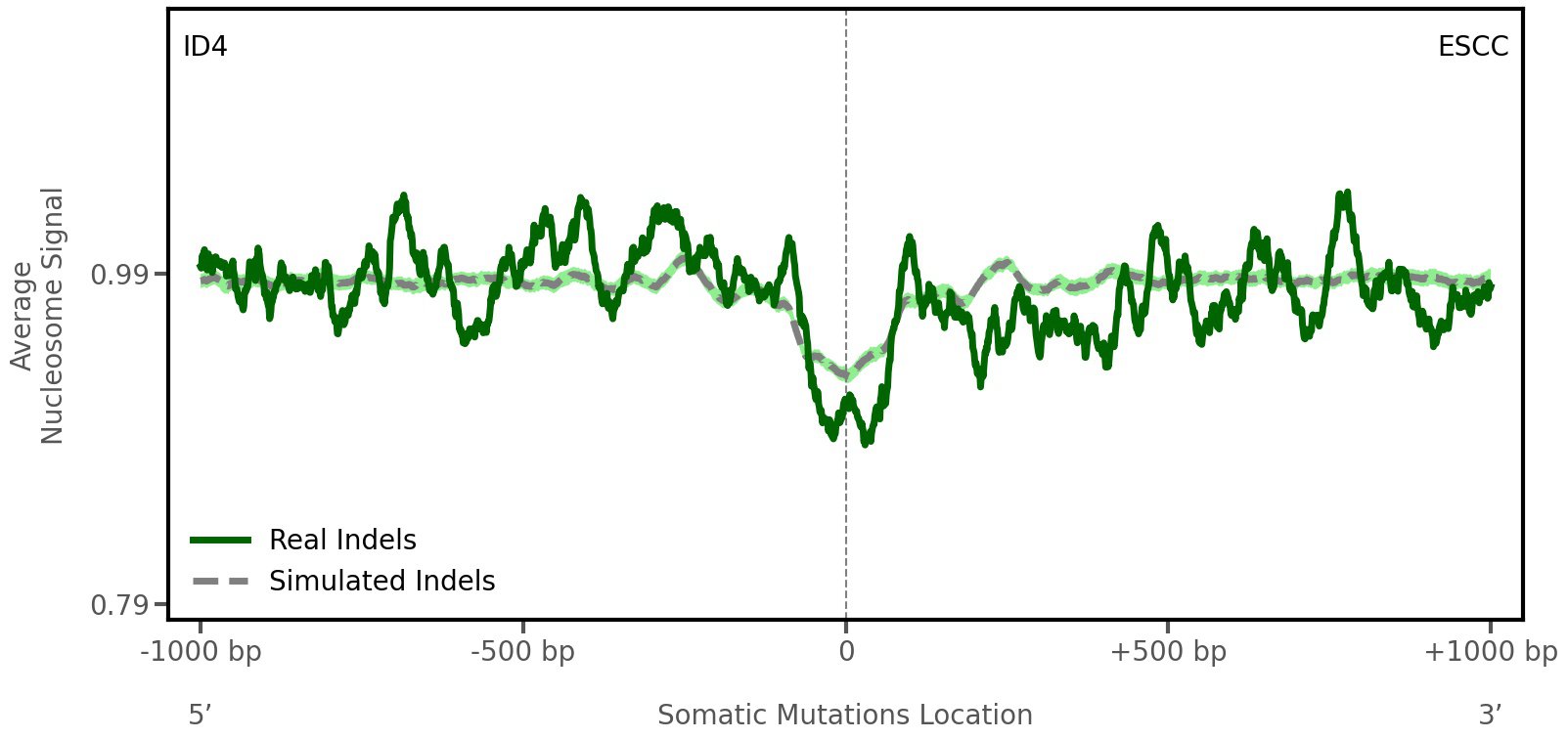
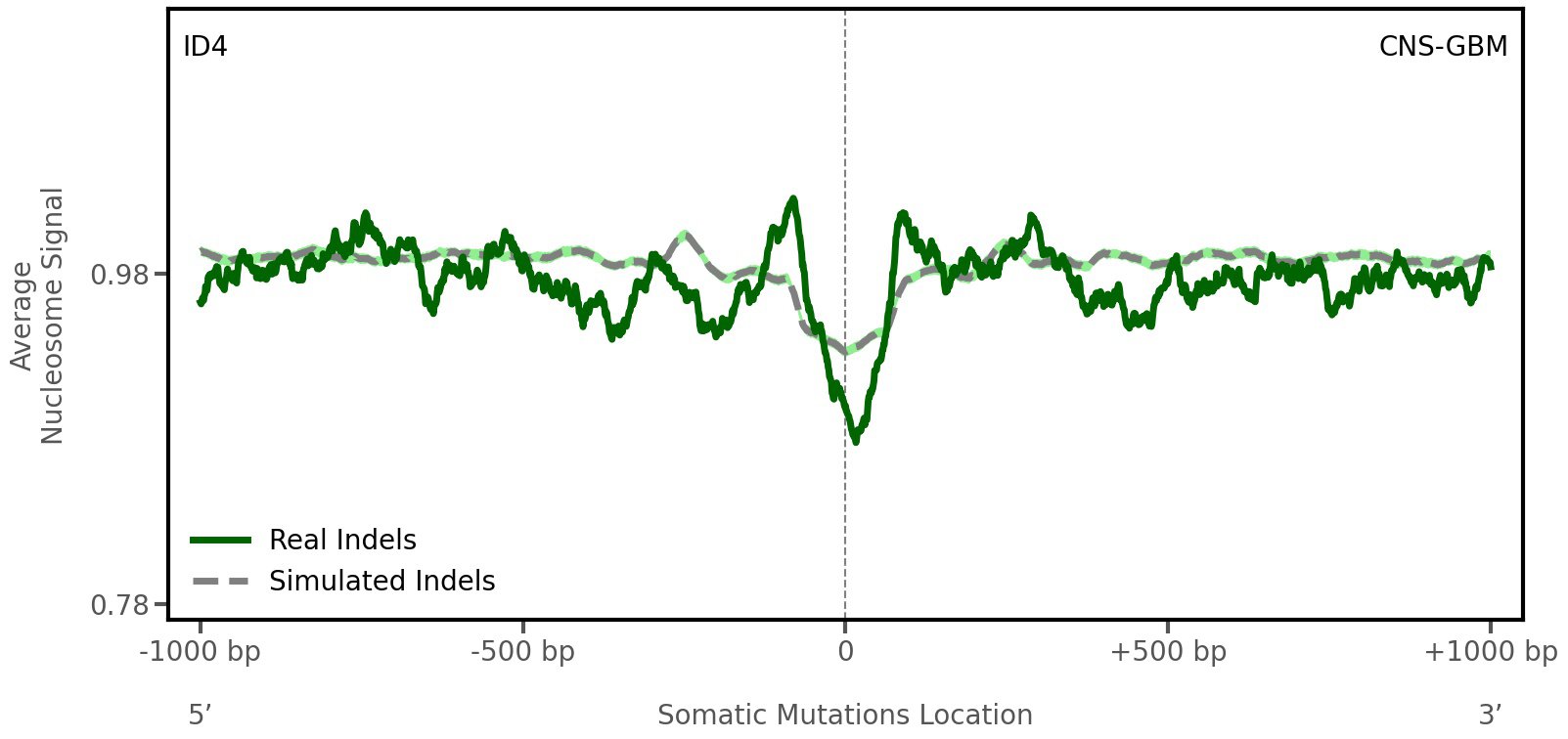
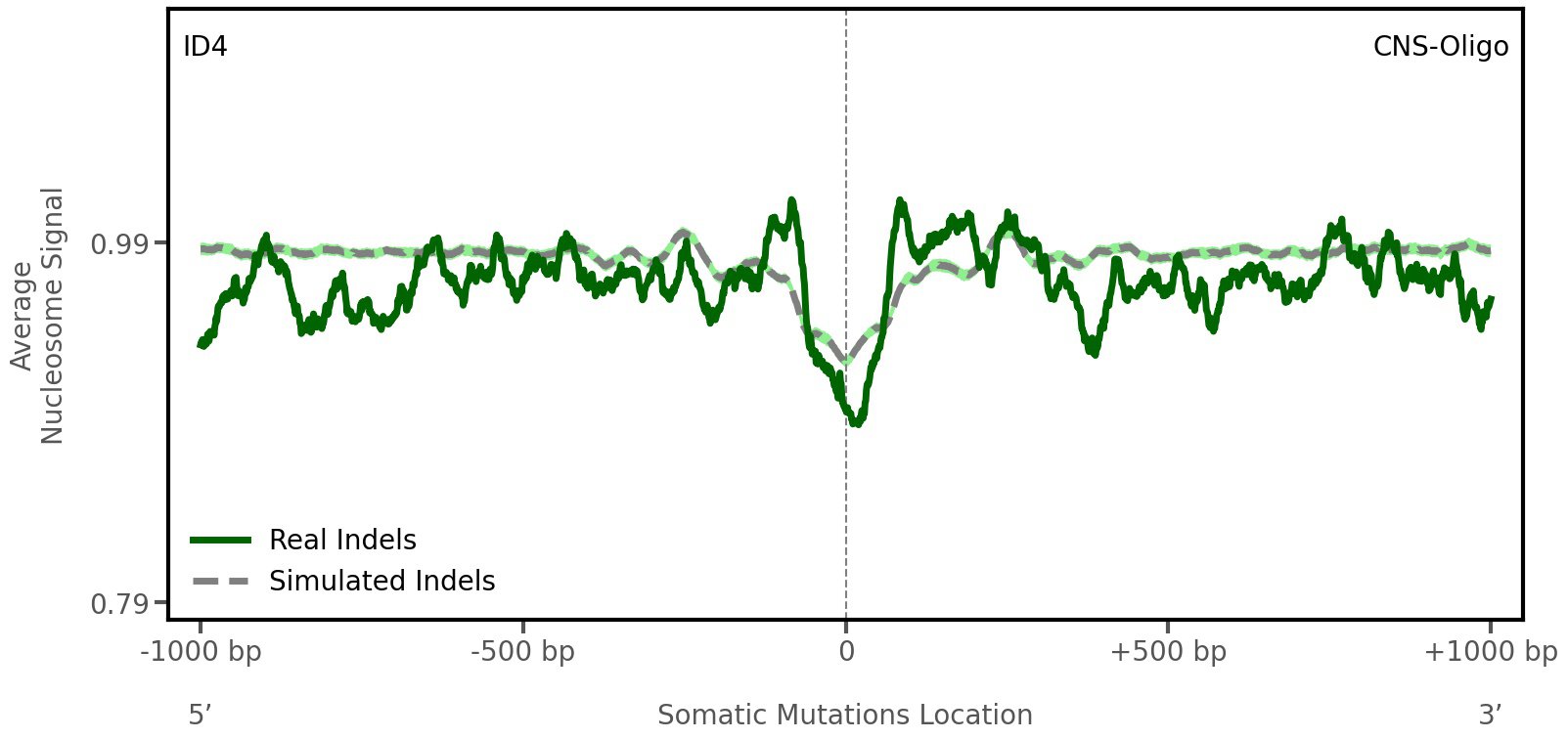
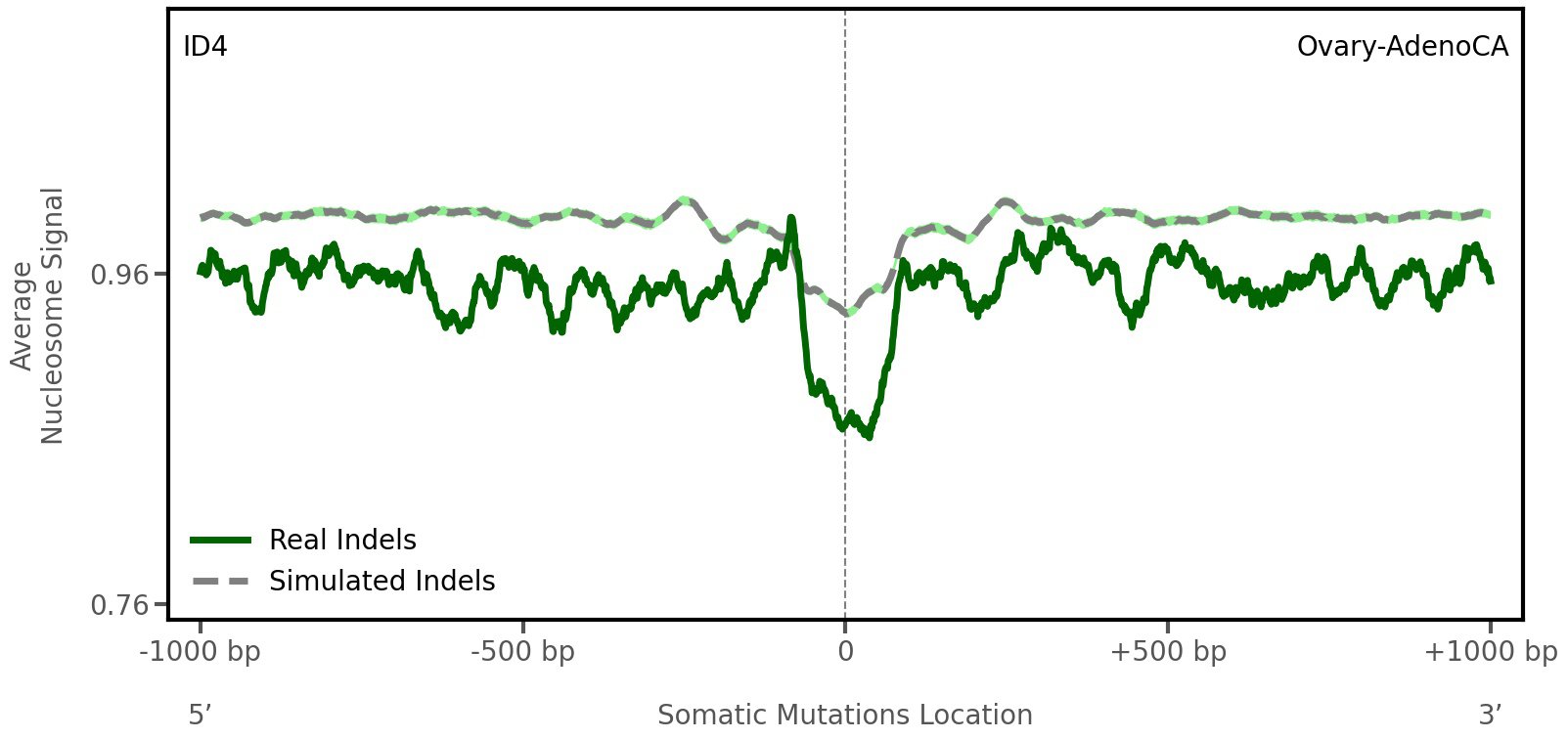
Nucleosome occupancy
Average nucleosome signal along a 2 kilobase window centred at the somatic mutation (dashed vertical line). The solid blue line shows the average nucleosome signal for real mutations, whereas the dashed line shows the average nucleosome signal for simulated somatic mutations. A higher signal reflects a higher propensity for nucleosome occupancy.
CTCF occupancy
Topography analysis could not be performed for CTCF occupancy as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
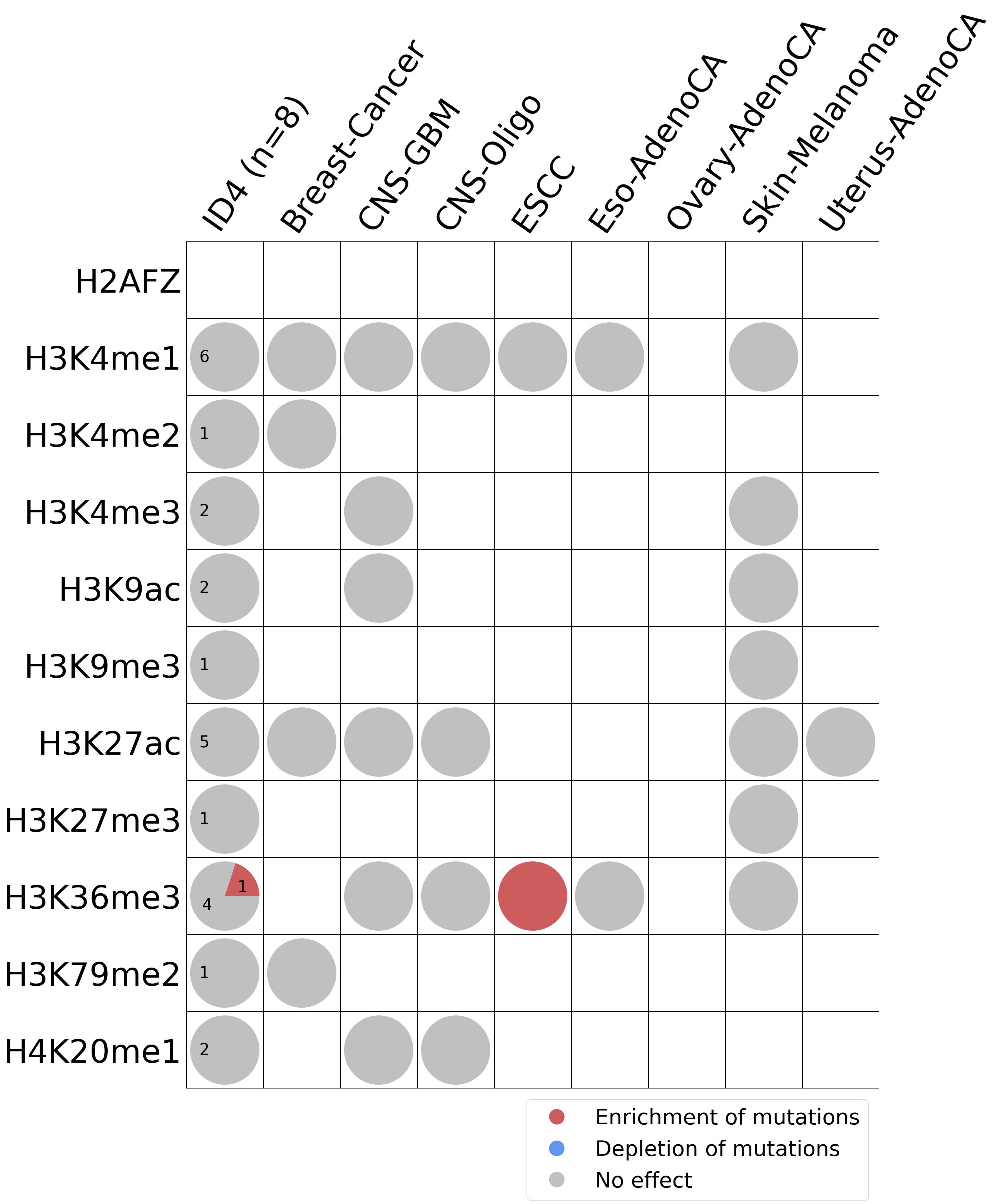
Histone modifications
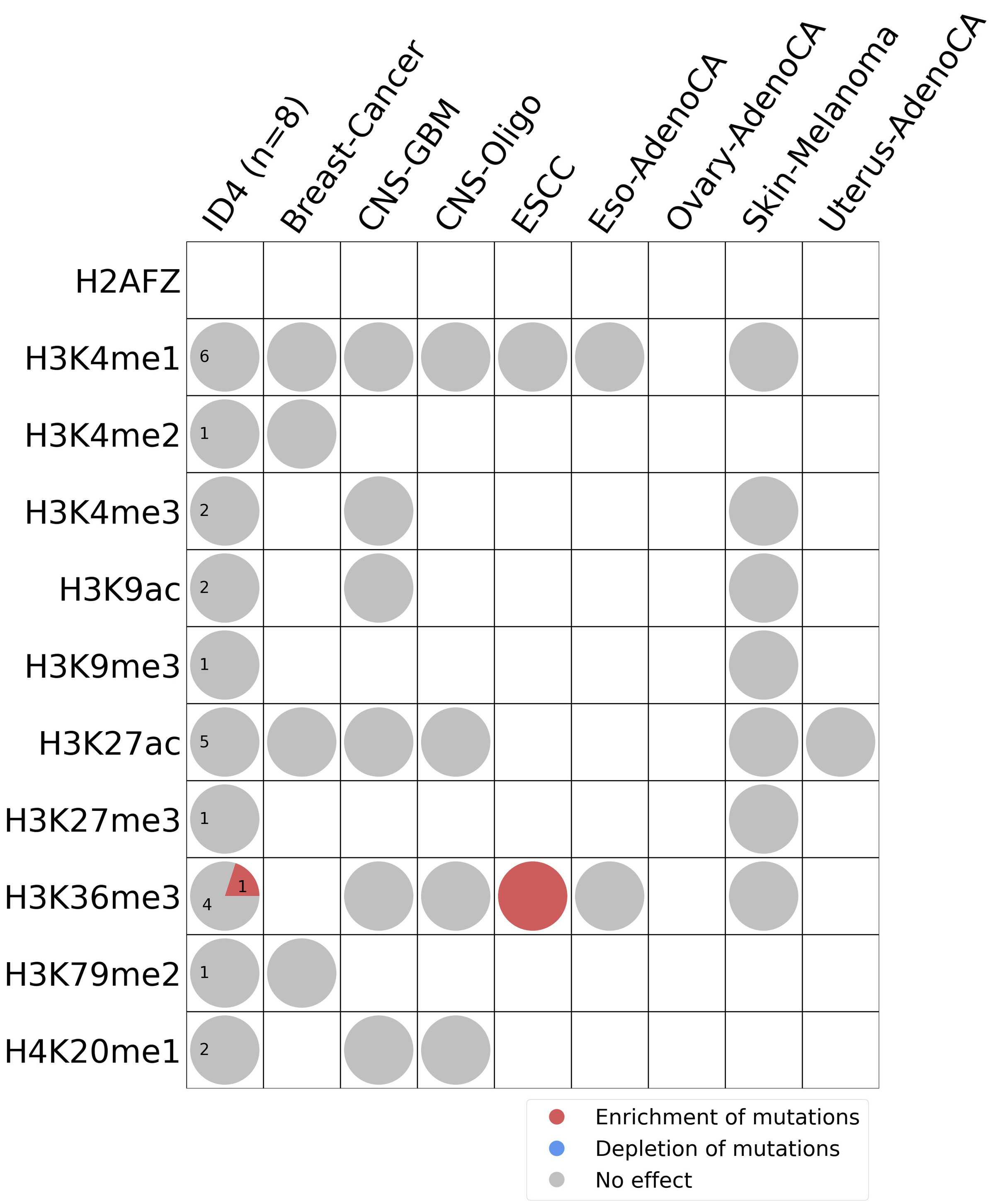
This plot shows the associations between mutational signatures and histone marks. Pie charts display the number of cancer types that are either enriched, depleted, or have no statistical effect for a given mutational signature and a specific histone mark. Differential changes are calculated by statistically comparing the of average signals between real and simulated mutations using a 100 base window centered at the somatic mutation. Enrichments reflect at least 5% statistically significant increases in real signal when compared to simulated signal. Depletions reflect at least 5% statistically significant decreases in real signal when compared to simulated signal. Statistical significance is determined based on false-discovery rate corrected p-value below 0.05.
(i) H2AFZ, a replication-independent member of the histone H2A family that renders chromatin accessible at enhancers and promoters and regulates transcriptional activation and repression; (ii) H3K4me1, histone mark often associated with enhancer activity; (iii) H3K4me2, a histone post-translational modification enriched in cis-regulatory regions, including both enhancers and promoters; (iv) H3K4me3, post-translational modification enriched in active promoters near transcription start sites; (v) H3K9ac, associated with active gene promoters and active transcription; (vi) H3K9me3, typical mark of constitutive heterochromatin; (vii) H3K27ac, histone modification generally contained at nucleosomes flanking enhancers; (viii) H3K27me3, repressive, associated with silent genes; (ix) H3K36me3, associated with transcribed regions and playing a role in regulating DNA damage repair; (x) H3K79me2, detected in the transcribed regions of active genes; and (xi) H4K20me1, found in gene promoters and associated with gene transcriptional elongation and transcription activation.
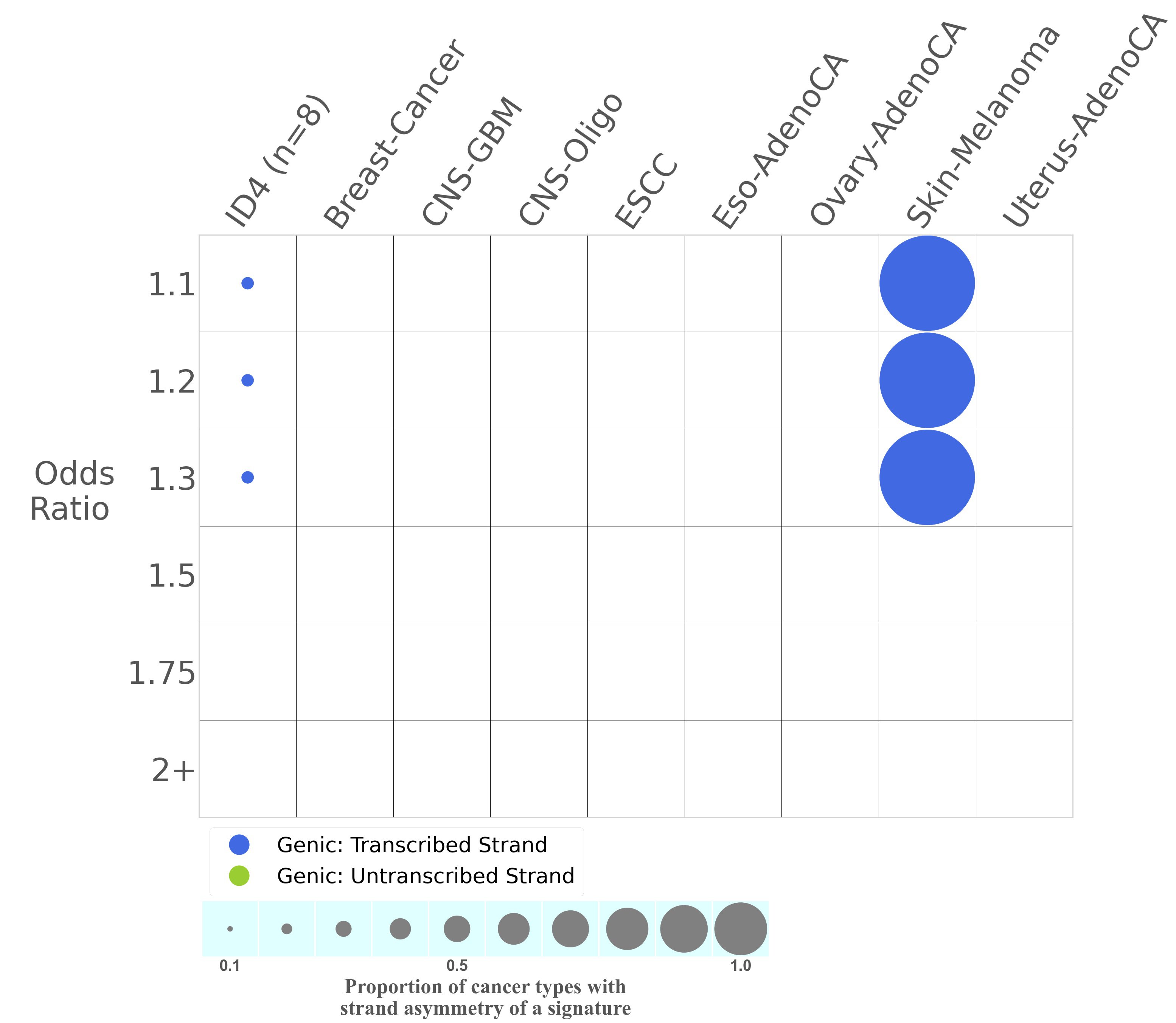
Transcriptional strand asymmetry
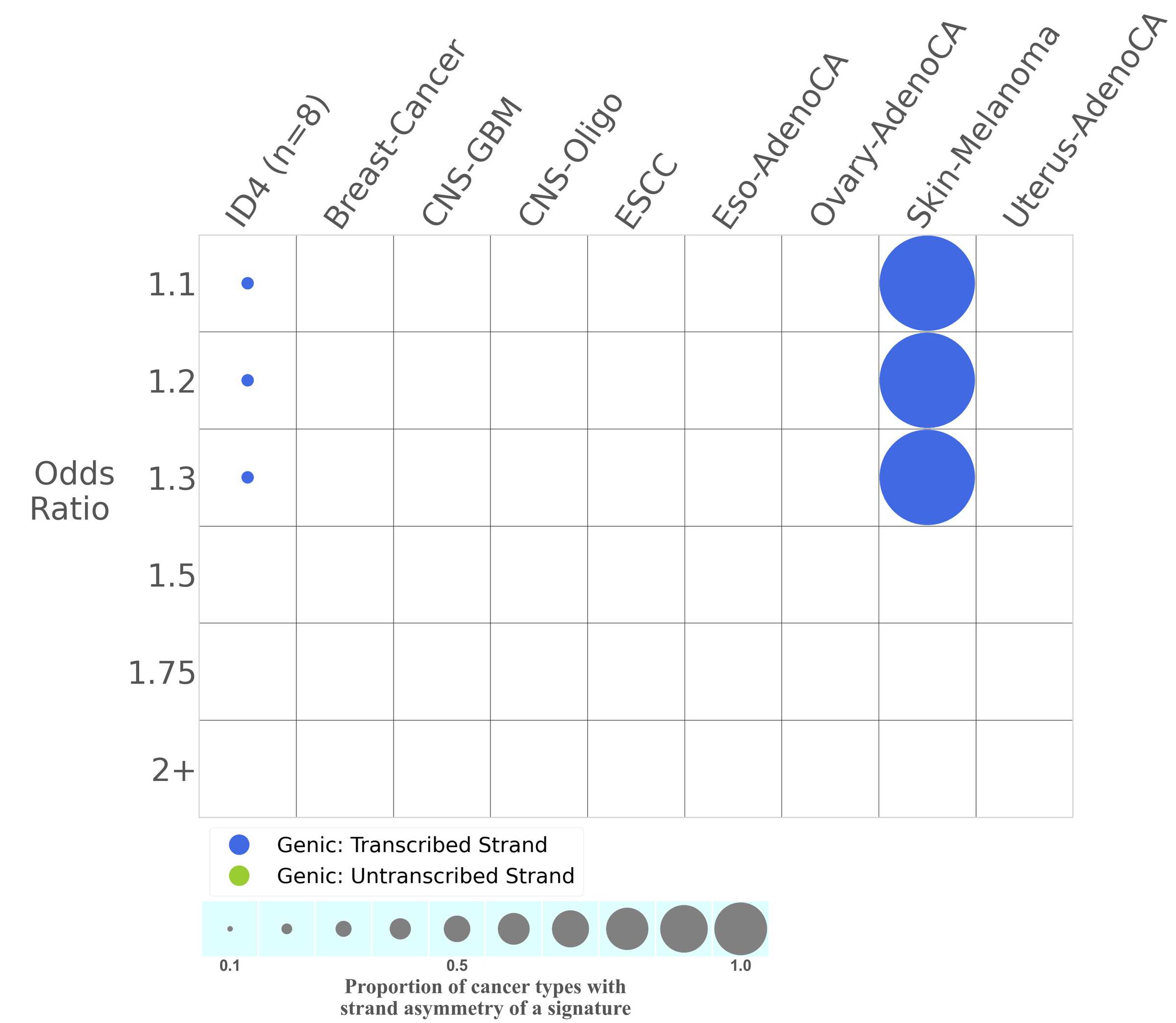
Mutational signatures exhibit asymmetric number of mutations due to either one of the DNA strands being preferentially repaired or one of the DNA strands having a higher propensity for being damaged. One common example of strand asymmetry is transcription-strand asymmetry which can be due to the activity of transcription-coupled nucleotide excision repair (TC-NER) or transcription-coupled damage amongst others.
The upper bar plot represents the percentage of real mutations in transcribed and untranscribed strands averaged across the human genome as well as all examined samples in 96 mutational context.
In the lower circle plot, the circles are filled with the colour of the significant strand when there is an odds ratio of at least 1.1 with statistical significance. The first row of this plot displays the transcriptional strand asymmetry across all cancer types, while the remaining rows present transcriptional strand asymmetry for each cancer type.
Each mutation attributed to the mutational signature is annotated as either on the transcribed strand, untranscribed strand, or non-transcribed strand.
The odds ratio is defined as the real mutations ratio divided by the simulated mutations ratio, where each ratio is calculated using the number of mutations on transcribed and untranscribed strands. And the number of real mutations on transcribed and untranscribed strands must be statistically significant with respect to the average number of simulated mutations on transcribed and untranscribed strands.
The strand with the higher number of real mutations defines the numerator of both the real mutations ratio and the simulated mutations ratio.
The strand with the lower number of real mutations defines the denominator of both the real mutations ratio and the simulated mutations ratio.
Genic and intergenic regions
Topography analysis could not be performed for genic and intergenic region asymmetry as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Replicational strand asymmetry
Topography analysis could not be performed for replicational strand asymmetry as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.