Mutational Signatures (v3.6 - May 2026)
ID18 · GRCh37 · COSMIC v104
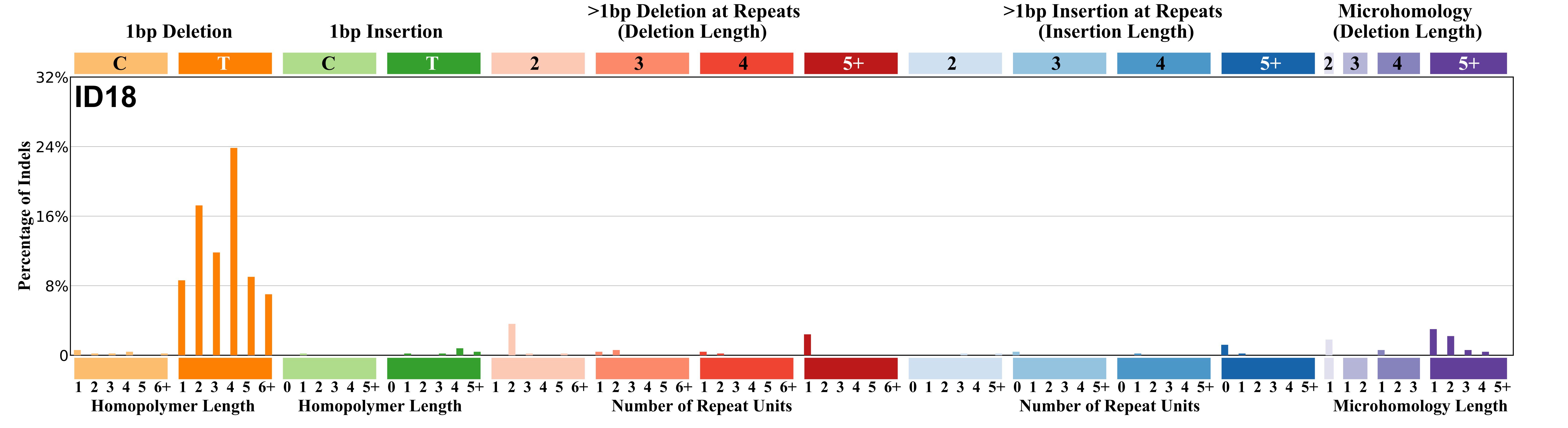
Mutational profile
The height of each mutational profile bar represents the proportion of one ID mutation type among all ID mutation types in the signature. Although there is no single intuitive and naturally constrained set of ID mutation types (as there arguably are for SBSs and DBSs), an 83 subclass categorisation of ID mutations was designed.
The 83 ID classification incorporates the prior knowledge that IDs commonly have sizes of 1-10 bps, that both insertions and deletions exist, that IDs of C and T occur at different rates, that IDs preferentially occur at repetitive elements, that the length of the repeat unit may influence the likelihood of an ID occurring, that the number of repeat units in a repeat stretch may influence the likelihood of an ID occurring, that IDs are also fostered in some instances by overlapping sequence microhomologies at the ID boundaries and that different mutational processes may, in principle, be differently influenced by these features. We therefore designed an 83 subclass categorisation of IDs that allows some exploration of all the above possibilities, while constraining the number of categories in order to accommodate the relatively small numbers of IDs (compared to substitutions) found in most genomes. This classification categorises IDs of lengths from 1bp to >5bp, for 1bp IDs classifies them as T or C and the number of single base repeats they occur in from 0 to >5, categorises lengths of non-single base repeat units from 2bp to >5bp and the number of repeats from 1 to >5 and size of microhomology from 0bp to >5bp. We recognise that different classifications of IDs may be preferred by others. The ID mutation types are enumerated in the following Excel document.
Proposed aetiology
Exposure to E.coli bacteria carrying pks pathogenicity island, producing genotoxic compound colibactin.
Comments
Strong preference for adenines 5' of homopolymer containing the deletion.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Lee-Six et al. 2019 Nature / Pleguezuelos-Manzano et al. 2020 Nature / Boot et al. 2020 Genome Research | v3.1 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | Yes | Preference for A 5' of homopolymer | |
| Proposed aetiology | Mutational process | Support | |
| Colibactin exposure | Experimental confirmation | ||
| Experimental validation | Experimental study | Species | |
| Pleguezuelos-Manzano et al. 2020 Nature | Human | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Mainly found in colorectal cancers and normal colorectal epithelial cells, as well as in some samples derived from head and neck cancer, urinary tract cancer and oral squamous cell carcinoma.
Associated signatures
Associated with SBS88, also arising from colibactin exposure.
Replication timing
Topography analysis could not be performed for replication timing as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Nucleosome occupancy
Topography analysis could not be performed for nucleosome occupancy as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
CTCF occupancy
Topography analysis could not be performed for CTCF occupancy as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Histone modifications
Topography analysis could not be performed for histone modifications as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Transcriptional strand asymmetry
Topography analysis could not be performed for transcriptional strand asymmetry as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Genic and intergenic regions
Topography analysis could not be performed for genic and intergenic region asymmetry as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.
Replicational strand asymmetry
Topography analysis could not be performed for replicational strand asymmetry as the number of mutations satisfying our constraints was insufficient or this signature was not yet analysed.