Mutational Signatures (v3.6 - May 2026)
DBS16 · GRCh37 · COSMIC v104
Mutational profile
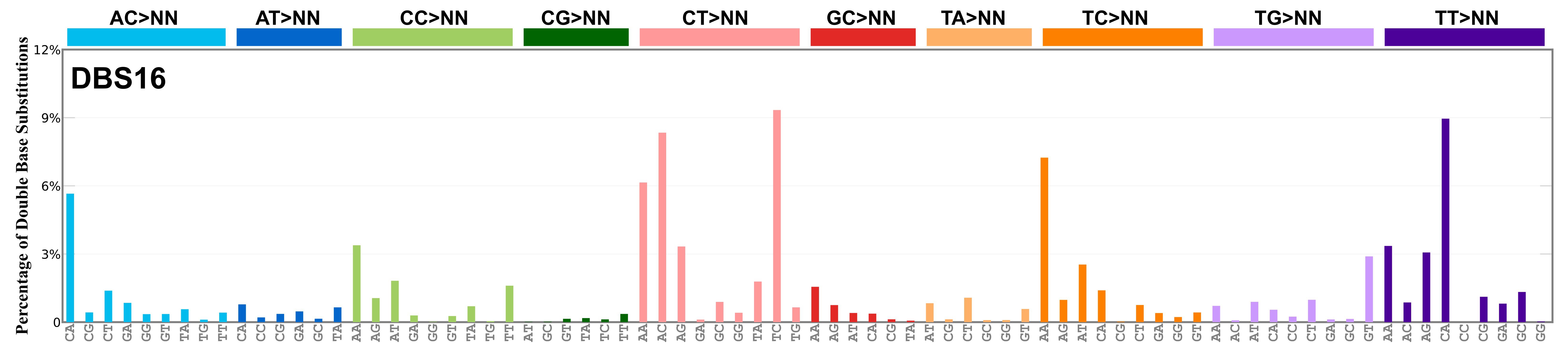
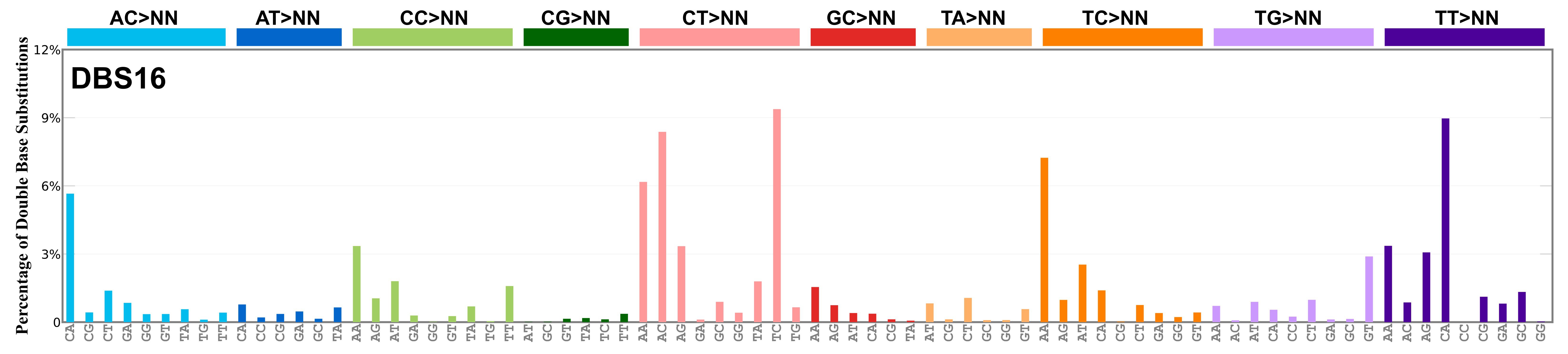
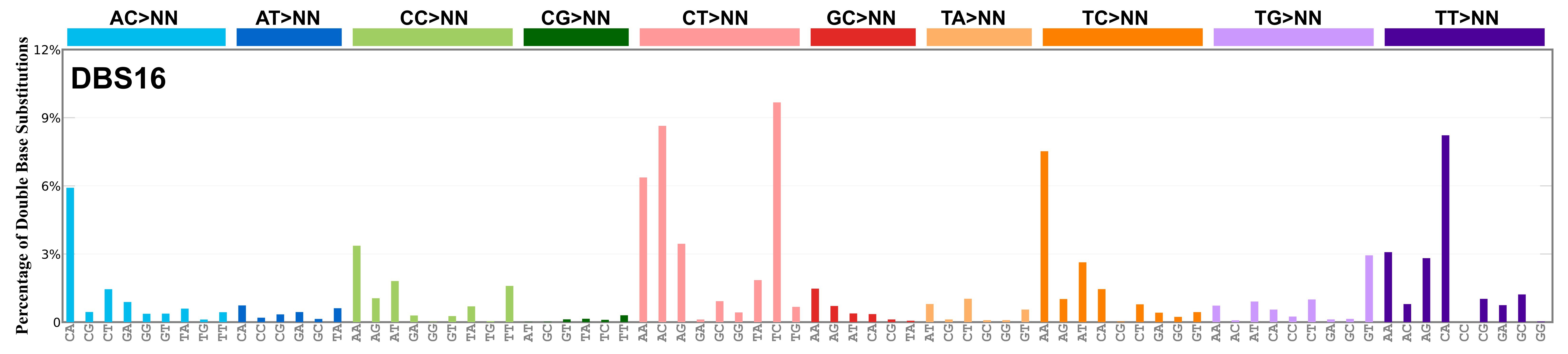
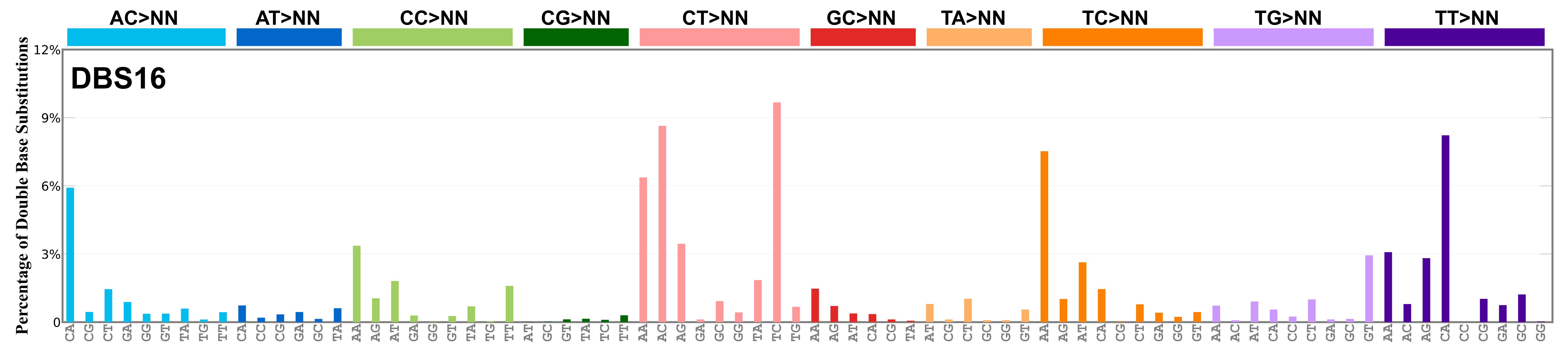
Proportion of a particular doublet base substitution (DBS) mutation type among all DBS mutation types in the signature is represented by the height of each bar. There are 78 strand-agnostic DBS mutation types.
The reason there are 78 strand-agnostic DBS mutation types is as follows. First, there are 4 x 4 = 16 possible source doublet bases. Of these, AT, TA, CG, and GC are their own reverse complement. We can represent the remaining 12 as 6 possible strand-agnostic doublets (e.g. AC represents both AC and its reverse complement, GT). Thus, there are 4+6=10 source doublet bases. Because they are their own reverse complements, AT, TA, CG, and GC can each be substituted by only 6 doublets . For example, AT can be substituted by 3 doublets starting with C: CA, CC, CG. But AT can be substituted by only 2 doublets starting with G: GA and GC. This is because the mutation from AT>GG is already represented by its reverse complement, AT>CC. Similarly AT can be substituted by only 1 doublet starting with T: TA. This is because AT>TC is represented by its reverse complement, AT>GA, and AT>TG is represented by AT>CA. For the remaining doublets, which are not their own reverse-complements, there are 3 x 3 = 9 possible DBS mutation types. Thus, in total there are 4 x 6 + 6 x 9 = 78 strand-agnostic DBS mutation types (see enumeration in the accompanying Excel document).
Proposed aetiology
Unknown.
Comments
Found in Degasperi et al. 2022 as DBS26. DBS16 includes a large contribution of CA>AC and TA>AT mutations, suggesting short inversion as its functional basis. Found in oesophagus and stomach, and it is possibly a variant or mixed version of RefSig DBS7. This signature might also be related to RefSig SBS17. Mutations are in cis, with the exception of TT>GG that is most likely caused by RefSig SBS17, in analogy with what is observed for RefSig DBS7 as noted in Degasperi et al. 2022.
Acceptance criteria
| Background | Identification study | First included in COSMIC | |
|---|---|---|---|
| Degasperi et al. 2022 Science | v3.4 | ||
| Identification | NGS technique | Different variant callers | Multiple sequencing centres |
| WGS | Yes | Yes | |
| Technical validation | Validated in orthogonal techniques | Replicated in additional studies | Extended context enrichment |
| Yes | Yes | - | |
| Proposed aetiology | Mutational process | Support | |
| Unknown | Unknown | ||
| Experimental validation | Experimental study | Species | |
| - | - | ||
Summary of the technical and experimental evidence available in the scientific literature regarding the validation of the mutational signature.
Tissue distribution
Common in upper gastrointestinal and a small fraction of lung cancers.